# Load Pandas
import pandas as pd # Import pandas
# Import the dataset specifying which Excel sheet name to load the data from
df_majors = pd.read_csv('https://raw.githubusercontent.com/ELSTE-Master/Data-Science/main/Data/Smith_glass_post_NYT_data_majors.csv')
df_traces = pd.read_csv('https://raw.githubusercontent.com/ELSTE-Master/Data-Science/main/Data/Smith_glass_post_NYT_data_traces.csv')Exploratory data analysis
How to start making data talk
October 22, 2025
Today’s objectives
- Last week: We learned how to handle data using
Pandas- Load, access, query, filter, sort and operate on data
- This week: You received / compiled / produced a new dataset
- Review common tasks / method to get accustomed to the dataset
- Understand its structure → what type of data does it contain
- Explore it visually → plotting with dedicated libraries
- Describe single variables → univariate analyses
- Assess coupled behaviours of two variables → bivariate analyses
- First attempt at modelling these behaviours → linear regressions
The dataset
Case study
- Campi Flegrei: Home to 1.5 million people
- Caldera-forming eruptions
- Campanian Ignimbrite (CI), ~39 ka ago
- Neapolitan Yellow Tuff (NYT), ~15 ka ago

Caldera-forming eruptions cycles
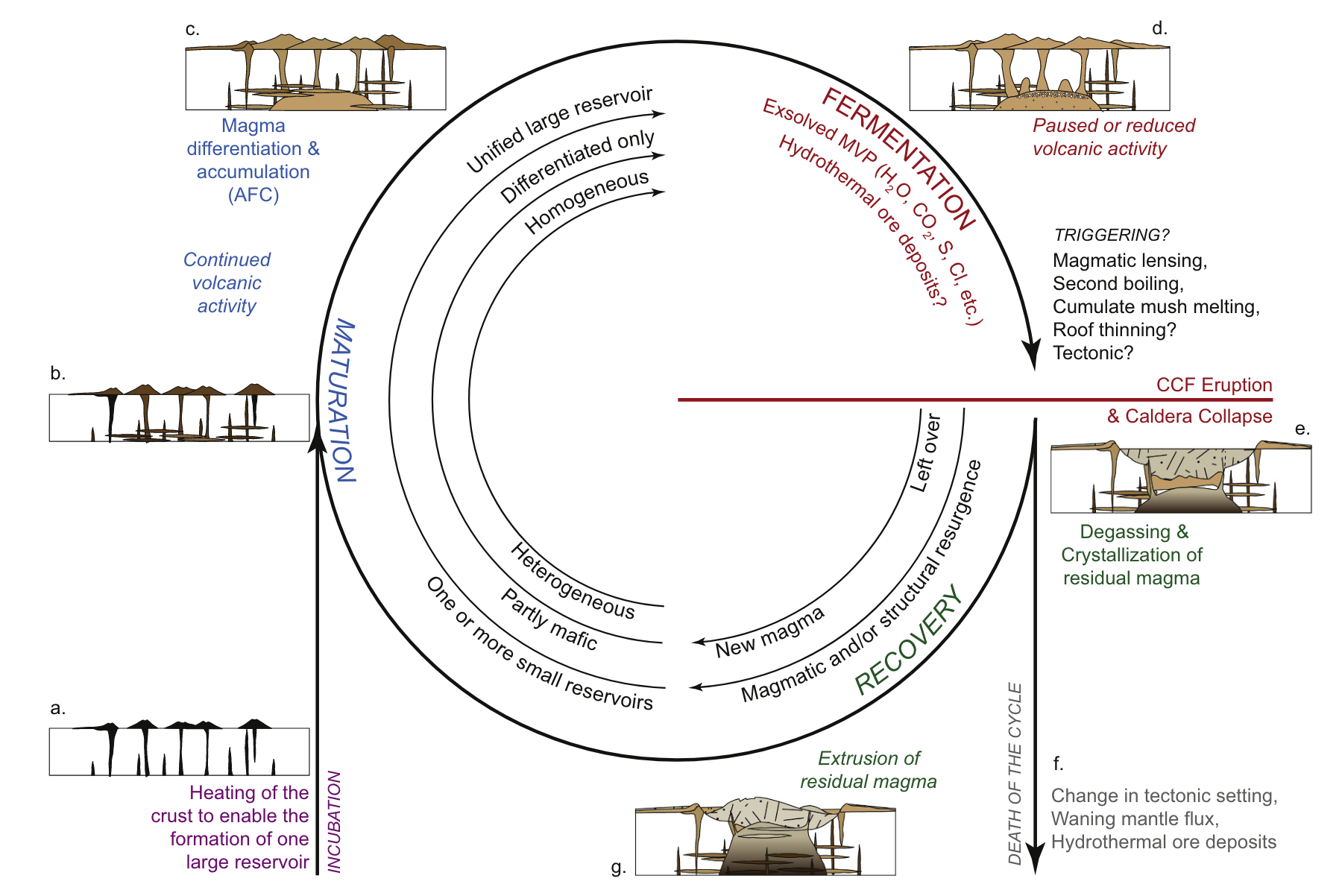
Bouvet de Maisoneuve et al. (2021)
Caldera cycles at Campi Flegrei
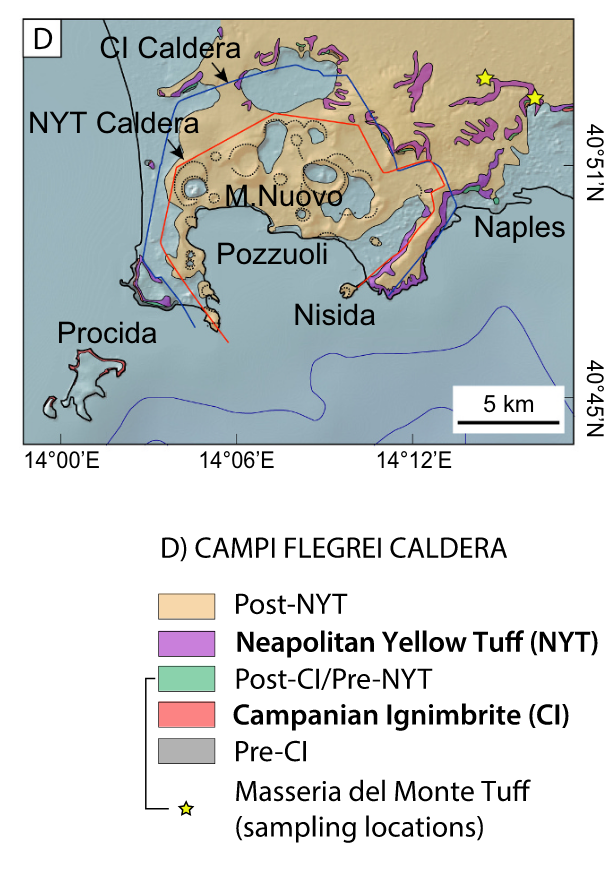
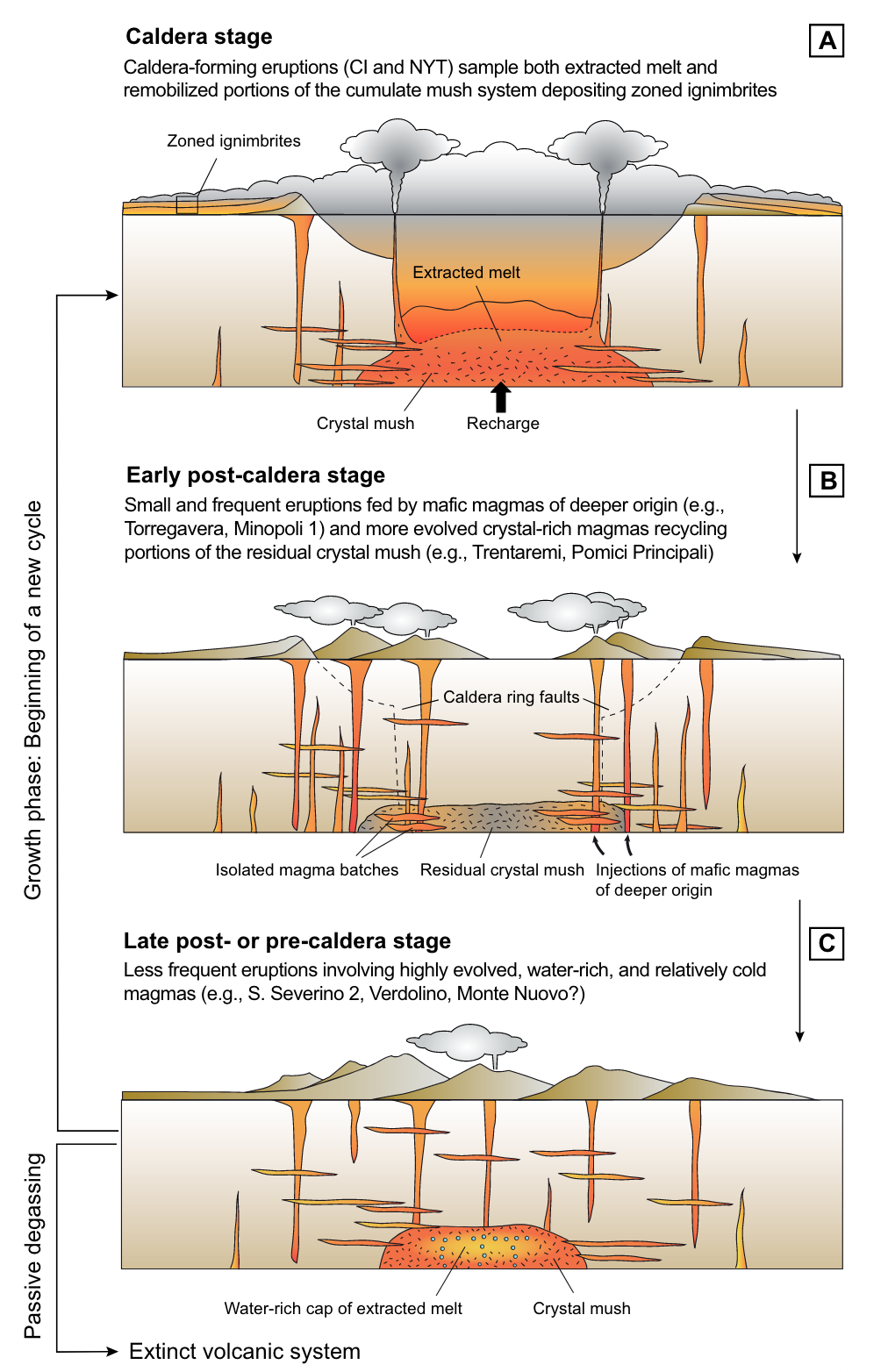
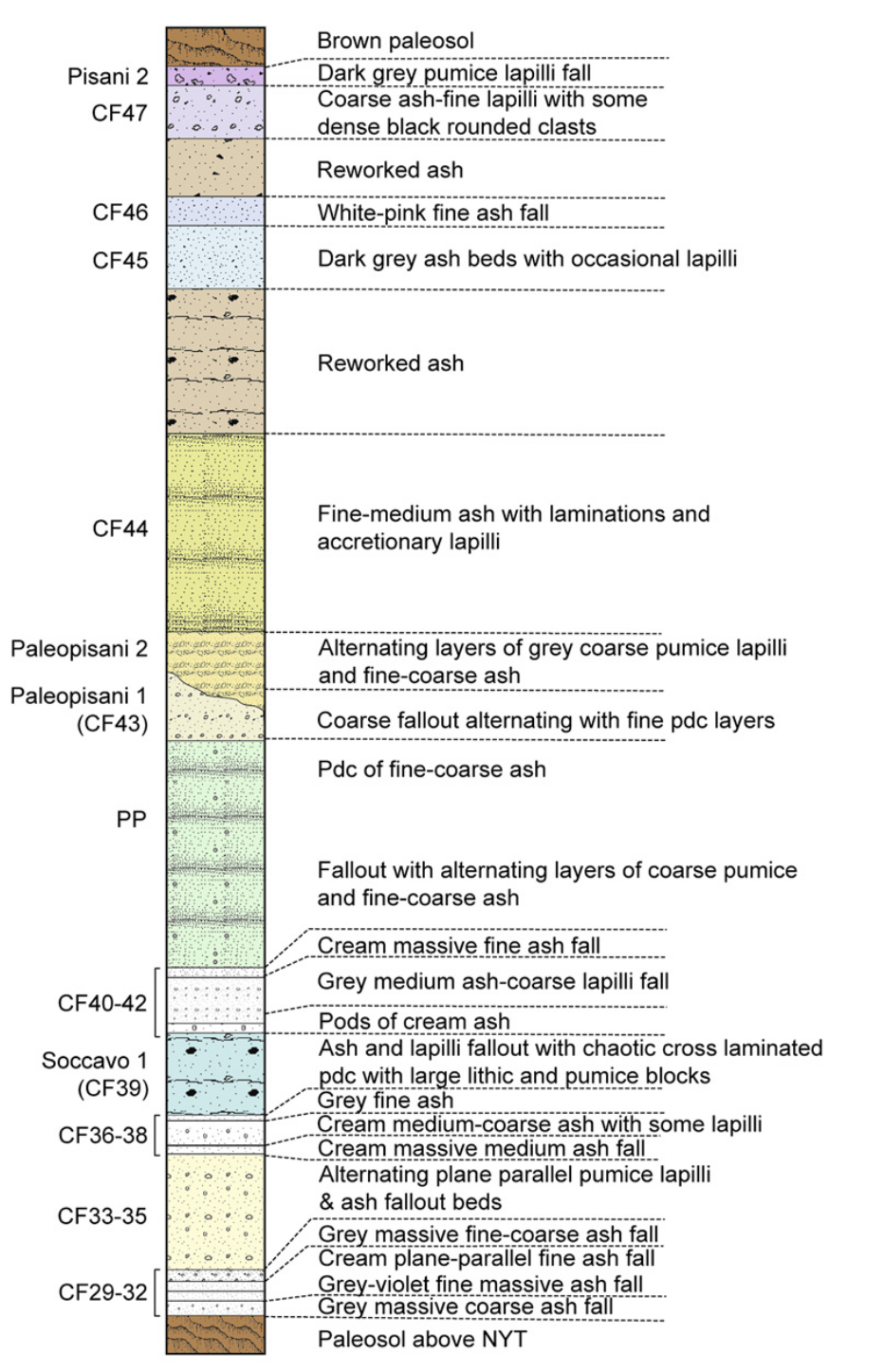
Dataset
Tephrostratigraphy and glass compositions of post-15 kyr Campi Flegrei eruptions
- Smith et al. (2011)
- Major and trace elements
Loading the dataset
- Import
pandas - Load an excel file using
pandas - Specify which sheet to read from using the
sheet_nameargument- Supp_majors →
df_majors - Supp_traces →
df_traces
- Supp_majors →
Inspecting the dataset
| Analysis no. | Strat. Pos. | Eruption | controlcode | Sample | Epoch | Crater size | Date of analysis | Si/bulk cps | SiO2* (EMP) | ... | Ho | Er | Tm | Yb | Lu | Hf | Ta | Pb | Th | U | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1915 | 63 | Astroni 7 | 1 | 79 | three-b | 20 | 100210am | 20.21 | 59.27 | ... | 1.11 | 3.26 | 0.47 | 2.80 | 0.43 | 7.84 | 2.96 | 60.93 | 35.02 | 9.20 |
| 1 | 1916 | 63 | Astroni 7 | 1 | 79 | three-b | 20 | 100210am | 11.92 | 59.27 | ... | 1.08 | 2.27 | 0.46 | 3.14 | 0.46 | 7.33 | 3.52 | 59.89 | 34.46 | 10.46 |
| 2 | 1917 | 63 | Astroni 7 | 1 | 79 | three-b | 20 | 100210am | 17.06 | 59.27 | ... | 1.25 | 3.69 | 0.61 | 3.51 | 0.63 | 8.43 | 3.05 | 49.87 | 29.22 | 8.73 |
| 3 | 1918 | 63 | Astroni 7 | 1 | 79 | three-b | 20 | 100210am | 24.52 | 59.27 | ... | 1.24 | 3.72 | 0.46 | 3.04 | 0.44 | 8.95 | 3.08 | 59.59 | 30.71 | 9.79 |
| 4 | 1919 | 63 | Astroni 7 | 1 | 79 | three-b | 20 | 100210am | 14.35 | 59.27 | ... | 1.08 | 2.68 | 0.46 | 2.79 | 0.41 | 7.24 | 2.67 | 60.70 | 32.13 | 9.01 |
5 rows × 37 columns
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 370 entries, 0 to 369
Data columns (total 37 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 Analysis no. 370 non-null int64
1 Strat. Pos. 370 non-null int64
2 Eruption 370 non-null object
3 controlcode 370 non-null int64
4 Sample 370 non-null object
5 Epoch 370 non-null object
6 Crater size 370 non-null int64
7 Date of analysis 370 non-null object
8 Si/bulk cps 370 non-null float64
9 SiO2* (EMP) 370 non-null float64
10 Sc 370 non-null float64
11 Rb 370 non-null int64
12 Sr 369 non-null float64
13 Y 370 non-null float64
14 Zr 370 non-null int64
15 Nb 370 non-null float64
16 Cs 370 non-null float64
17 Ba 370 non-null float64
18 La 370 non-null float64
19 Ce 370 non-null float64
20 Pr 370 non-null float64
21 Nd 370 non-null float64
22 Sm 370 non-null float64
23 Eu 370 non-null float64
24 Gd 370 non-null float64
25 Tb 370 non-null float64
26 Dy 370 non-null float64
27 Ho 370 non-null float64
28 Er 370 non-null float64
29 Tm 370 non-null float64
30 Yb 370 non-null float64
31 Lu 369 non-null float64
32 Hf 370 non-null float64
33 Ta 370 non-null float64
34 Pb 368 non-null float64
35 Th 370 non-null float64
36 U 370 non-null float64
dtypes: float64(27), int64(6), object(4)
memory usage: 107.1+ KBTypes of data
Basic types of data
int: Integer numbers → 0, 1, 2, …float: Decimal numbers → 1.1, 1.2, 1.3, …object: Strings → “Campi Flegrei”
A note on Bytes
intandfloatnumbers are followed by 8, 16, 32 or 64 → Byte → control how much memory a variable uses- A Bit is a binary digit → 0/1 → smallest possible unit of data
int8can store \(2^8\) digits = 256int16can store \(2^16\) digits = 65’563
Families of data
Numerical data
- Represent quantities or measurable values
- Quantitative
- Discrete: Earthquake counts
- Continuous: Temperature
- Stored as numerical values
- Operations → arithmetic
Categorical data
- Represent categories or groups of data
- Qualitative / semi-quantitative
- Nominal: No order → Landslide type
- Ordinal: Order → rank
- Stored as strings or integer
- Operations → counting, grouping
Families of data - cheat sheet
| Feature | Categorical Data | Numerical Data |
|---|---|---|
| Definition | Categories or groups of data | Quantities or measurable values |
| Nature | Qualitative (describes qualities) | Quantitative (describes amounts or measurements) |
| Data Type | Non-numeric (often text or labels) | Numeric (numbers only) |
| Examples | Eruption type | Element concentration |
| Possible Operations | Counting, grouping, mode | Arithmetic operations |
| Measurement Scale | Nominal or Ordinal | Interval or Ratio |
| Visualization Tools | Bar chart, Pie chart | Histogram, Box plot, Scatter plot |
| Subtypes | Nominal: Categories with no order (e.g., color); Ordinal: Categories with order (e.g., rank) | Discrete: Countable numbers (e.g., number of earthquakes); Continuous: Measurable values (e.g., temperature) |
| Examples of Statistical Summary | Frequency, mode | Mean, median, standard deviation |
| Storage Format | Strings or labels | Integers or floats |
Plotting data
Plotting libraries
There are two main plotting libraries:
matplotlib(and its modulepyplot): core of plotting in Pythonseaborn: based onmatplotlib, designed for statistical exploration, integration withpandas
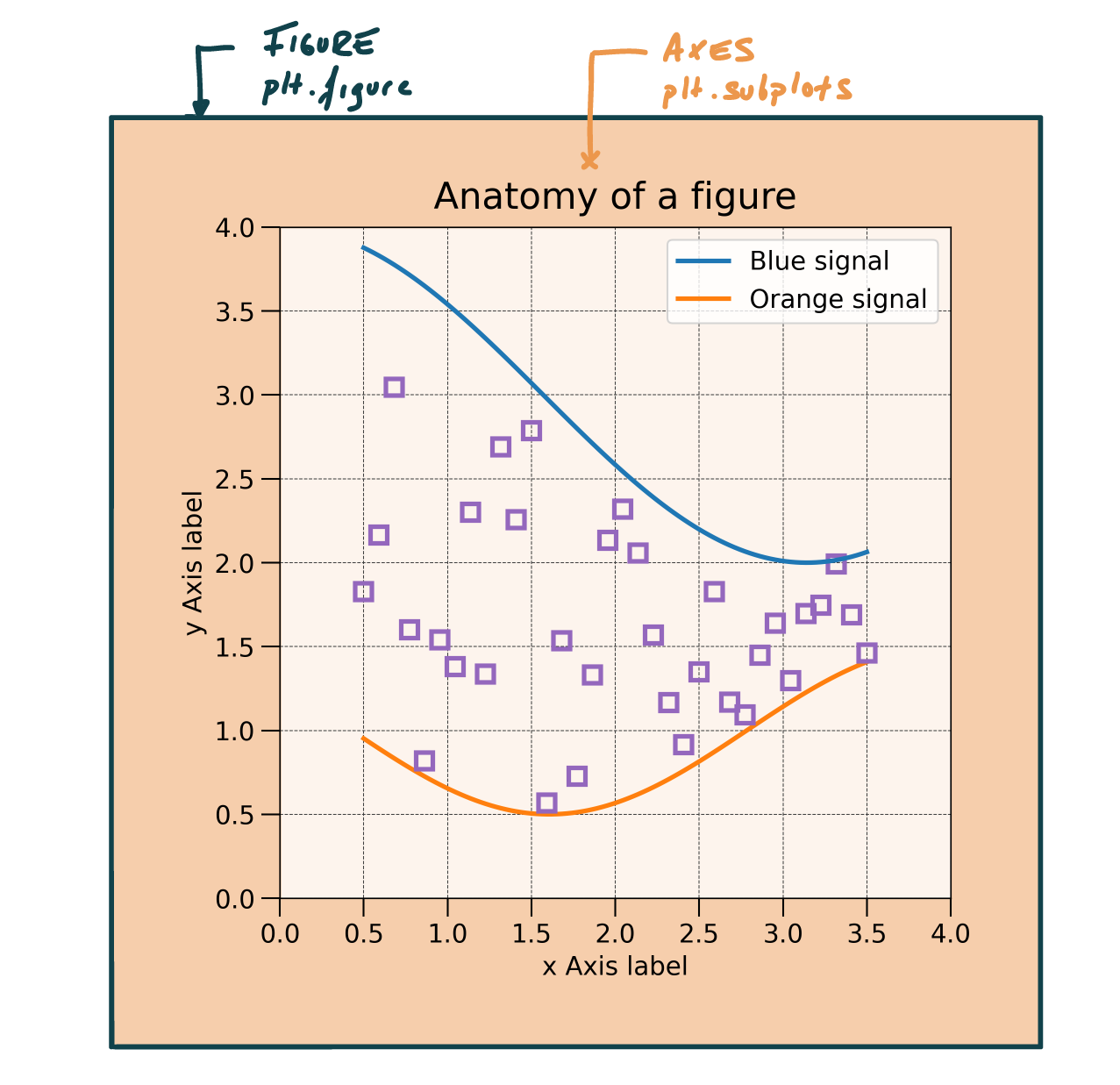
Anatomy of a figure
Anatomy of a axes
Axes: Where most of the magic occurs
| Function | Description |
|---|---|
ax.set_title |
Sets the title of the axes |
ax.set_xlabel |
Sets the label for the x-axis |
ax.set_ylabel |
Sets the label for the y-axis |
ax.legend |
Displays the legend |
ax.grid |
Shows grid lines |
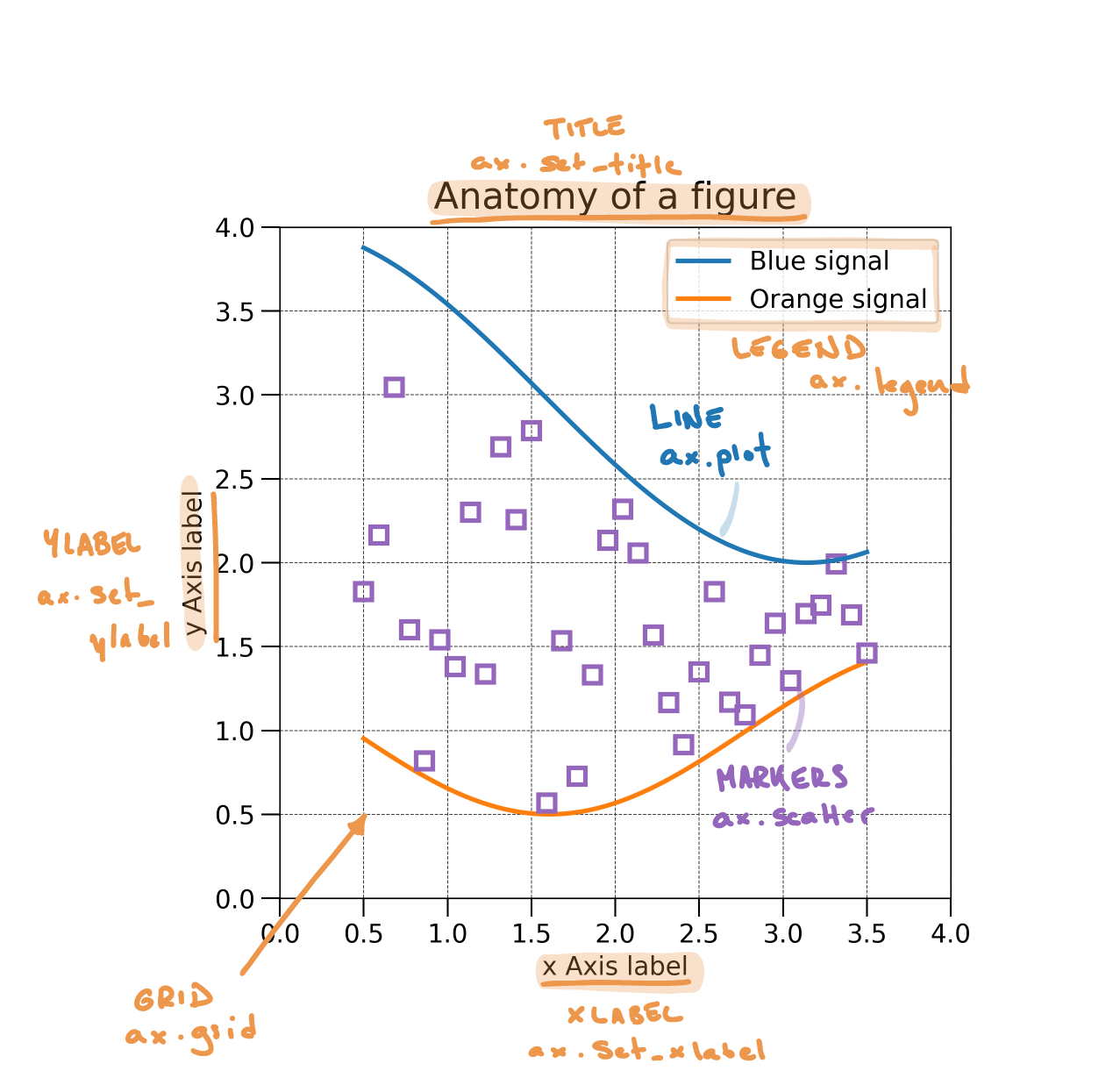
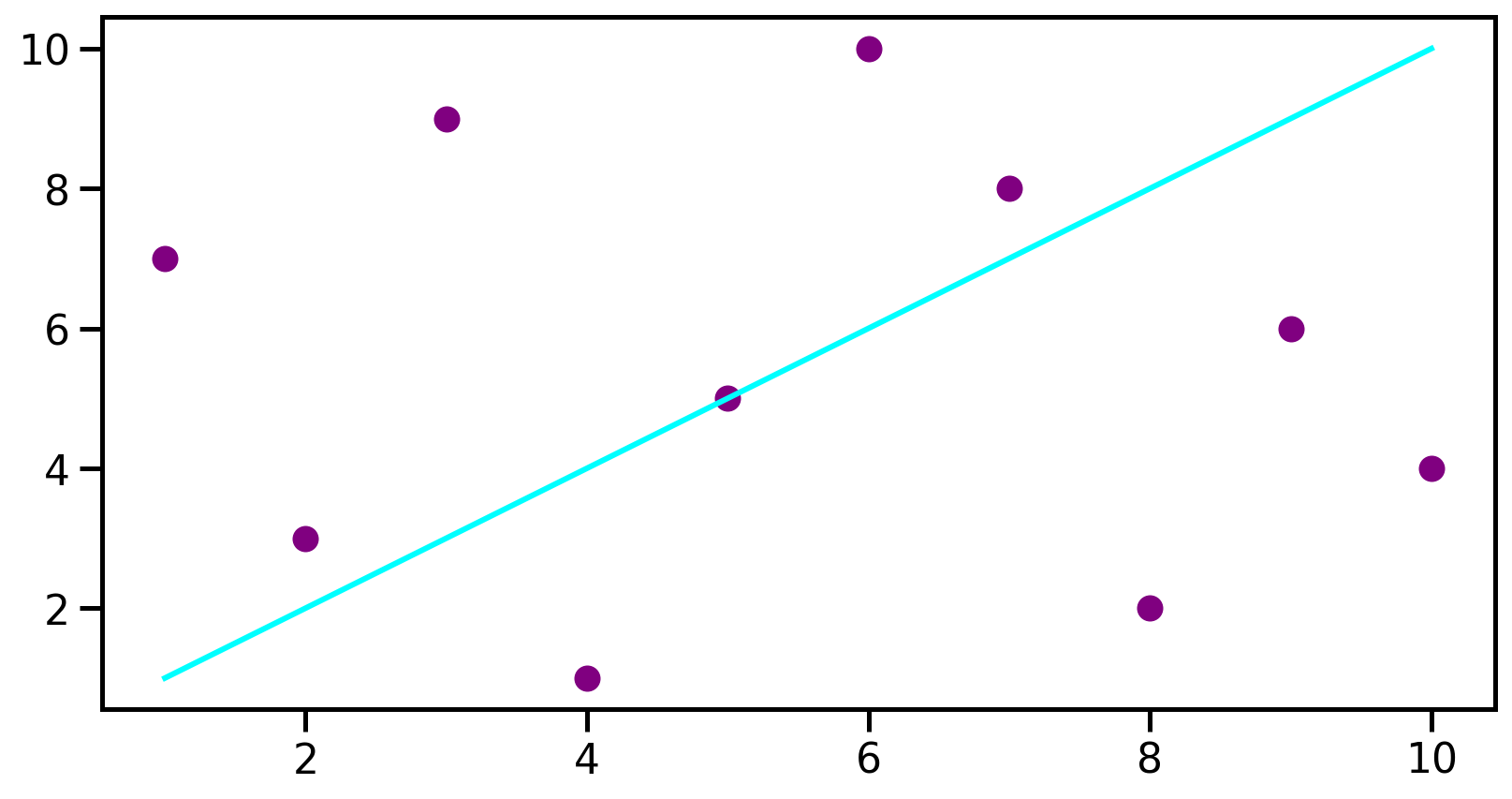
Plotting example
Plotting example
# Define some data
data1 = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
data2 = [7, 3, 9, 1, 5, 10, 8, 2, 6, 4]
# Set the figure and the axes
fig, ax = plt.subplots()
# Plot the data
ax.plot(data1, data1, color='aqua', label='Line')
ax.scatter(data1, data2, color='purple', label='scatter')
# Set labels
ax.set_xlabel('x Label')
ax.set_ylabel('y Label')
# Only to make it look pretty on the presentation
plt.show()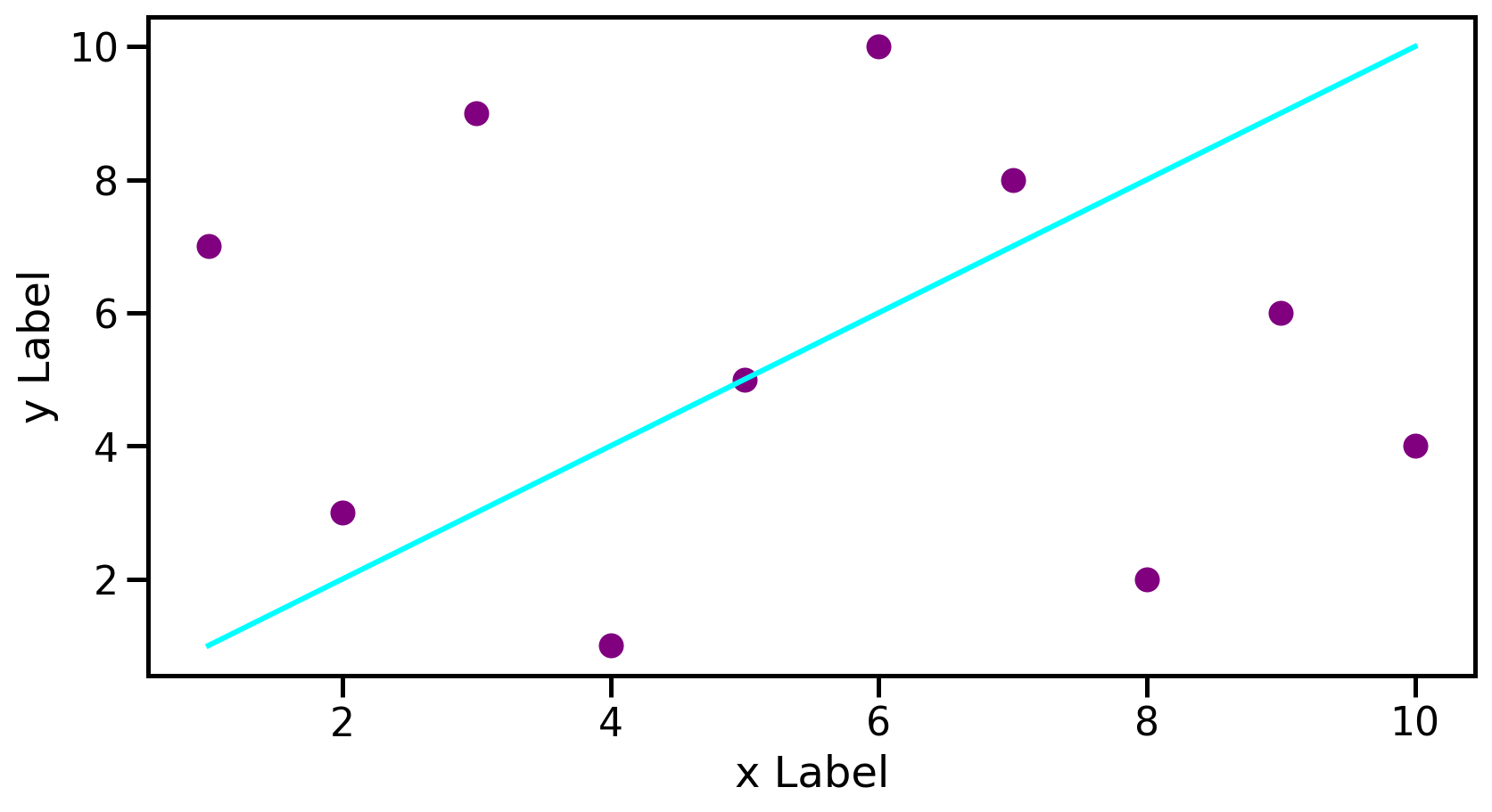
Seaborn
Why Seaborn?
Seaborn provides a high-level interface for drawing attractive and informative statistical graphics.
- Makes plotting
pandasDataFrames easy - Produces clean graphics
- Drawback: difficult customisation
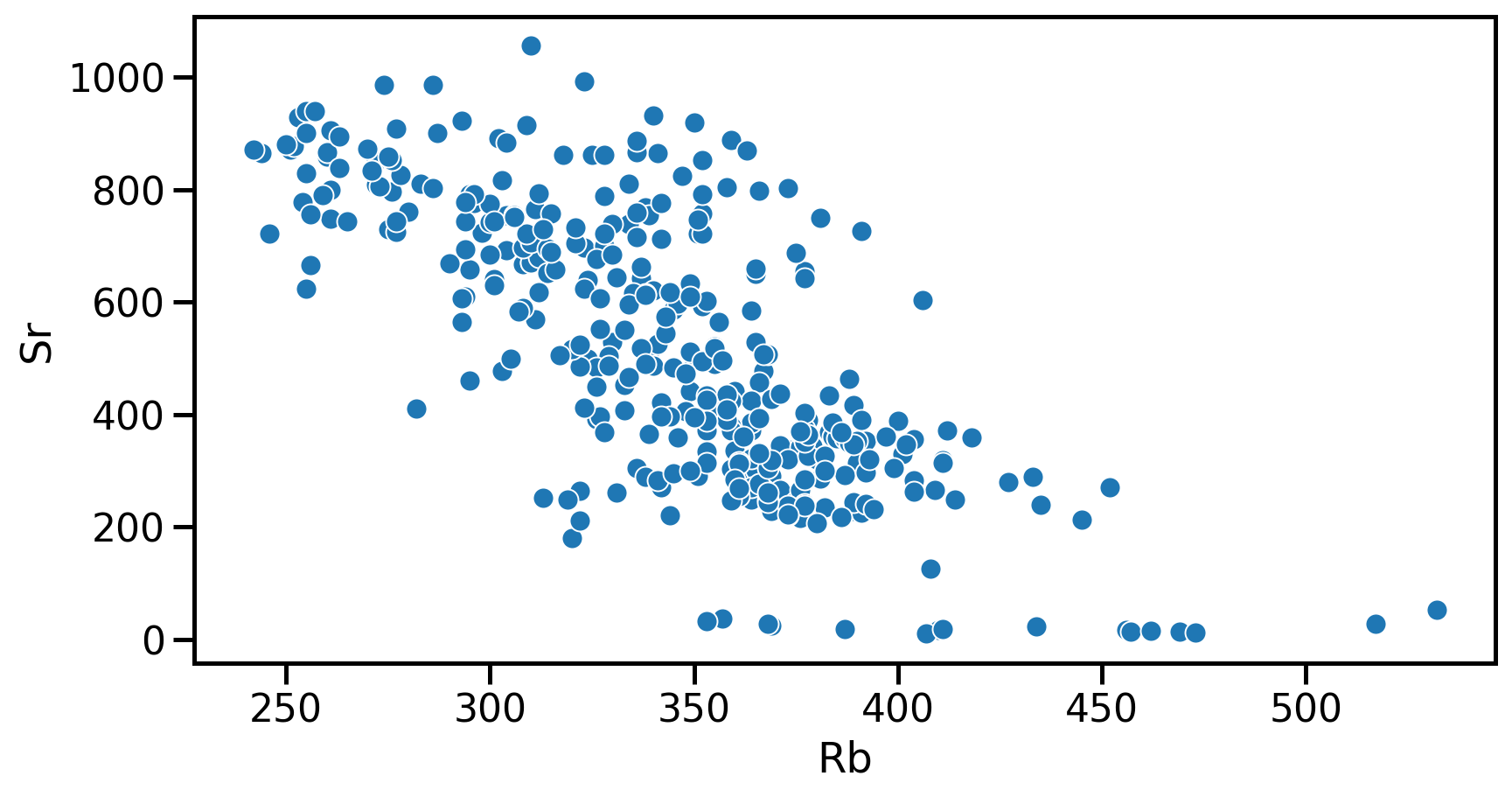
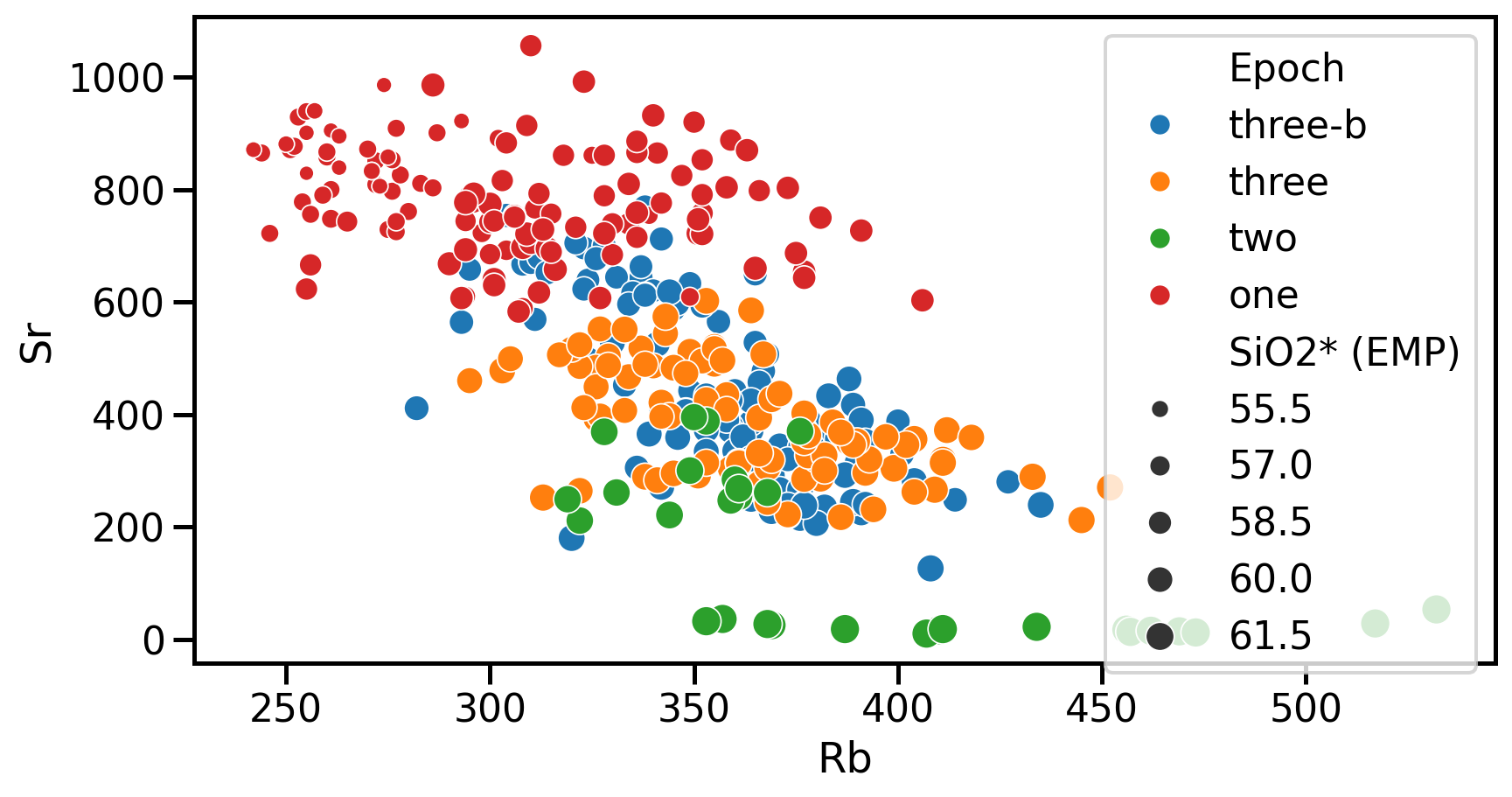
Seaborn example
Seaborn example
Your turn!
- Go to https://elste-master.github.io/Data-Science/
- Class 2 > Data exploration
- Explore the structure of the glass geochemistry or your dataset
- Get used to plotting functions
Descriptive statistics
Descriptive statistics
- Descriptive statistics → provide ways of capturing the properties of a given data set or sample.
PandasandSeaborn→ review metrics, tools, and strategies to summarize a dataset- Providing us with a quantitative basis to describe it and compare it to other datasets
Univariate analyses
Univariate analyses
- Objective capturing the properties of single variables at the time
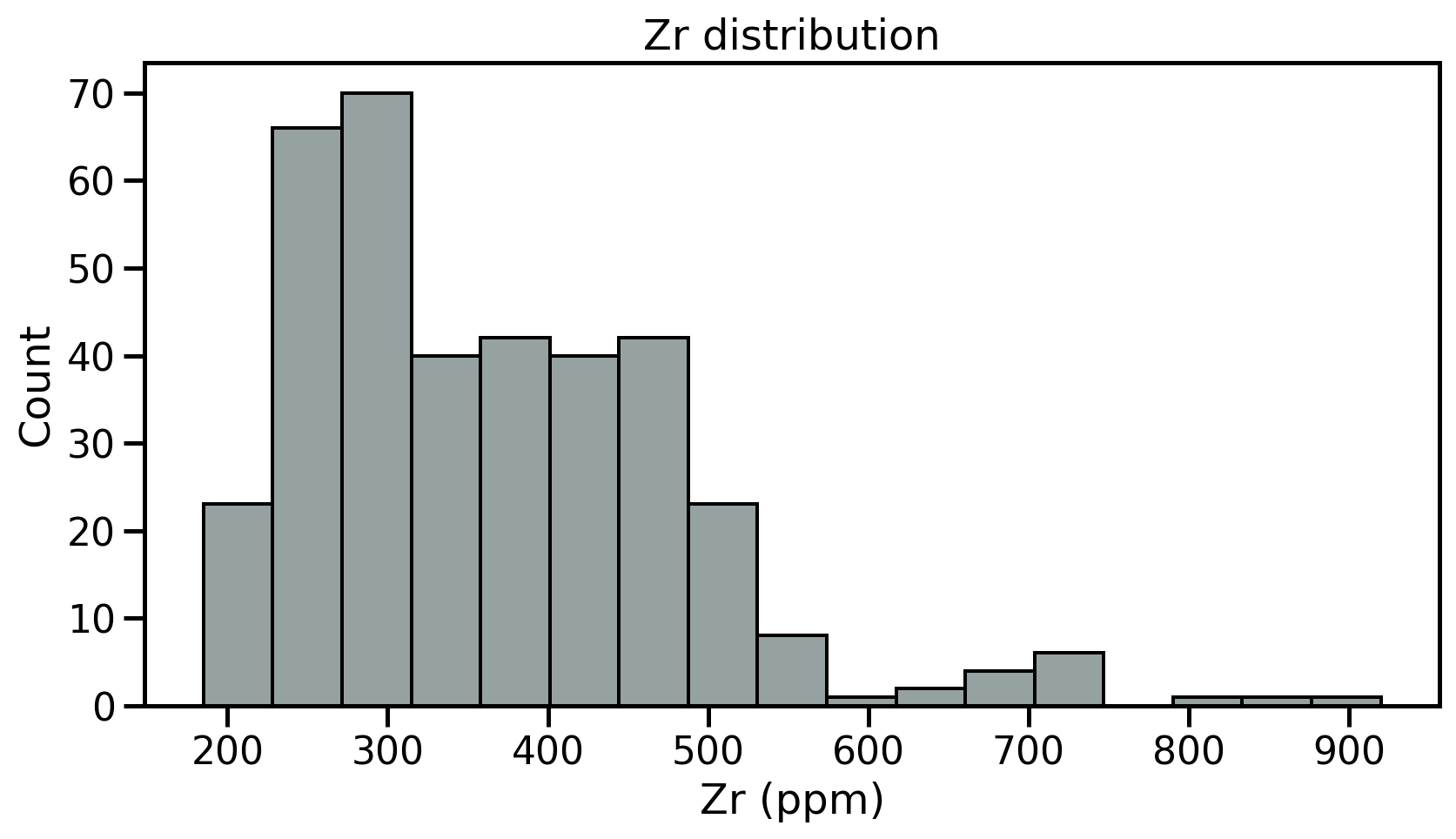
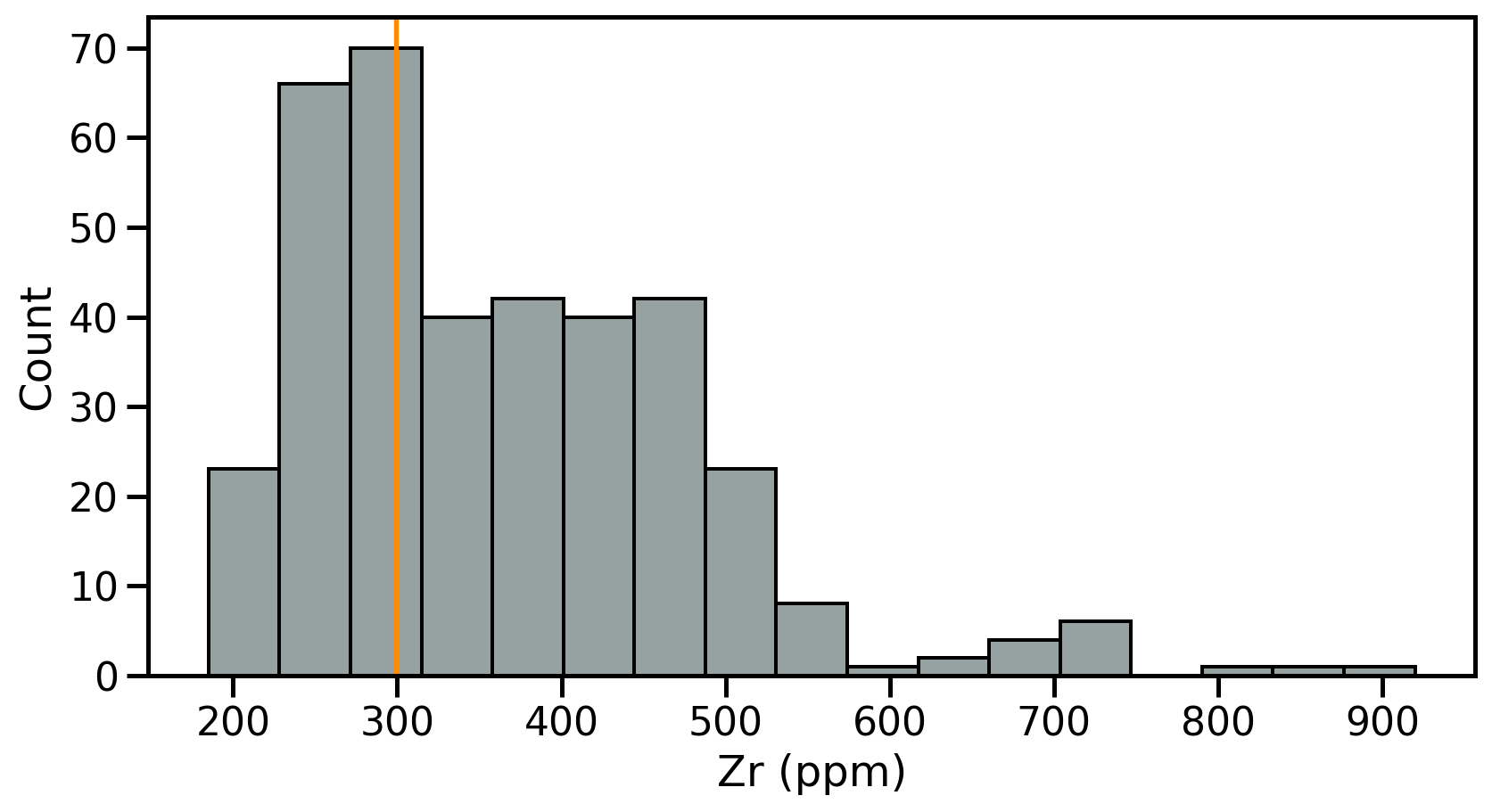
- Fist step: review the samples distribution using histograms
- Divides dataset into equal intervals → bins
- Counts the value in each bin
- Represents the distribution of data
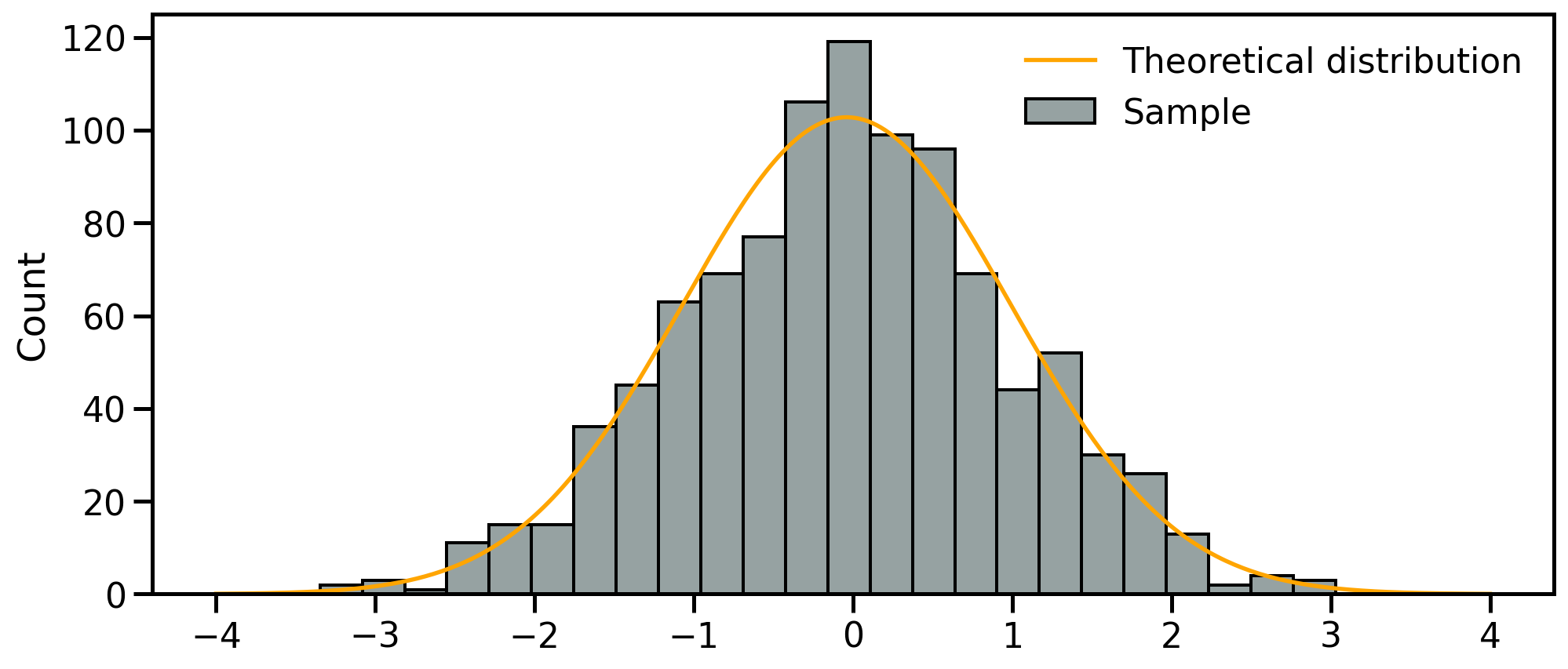
Univariate analyses
- Objective capturing the properties of single variables at the time
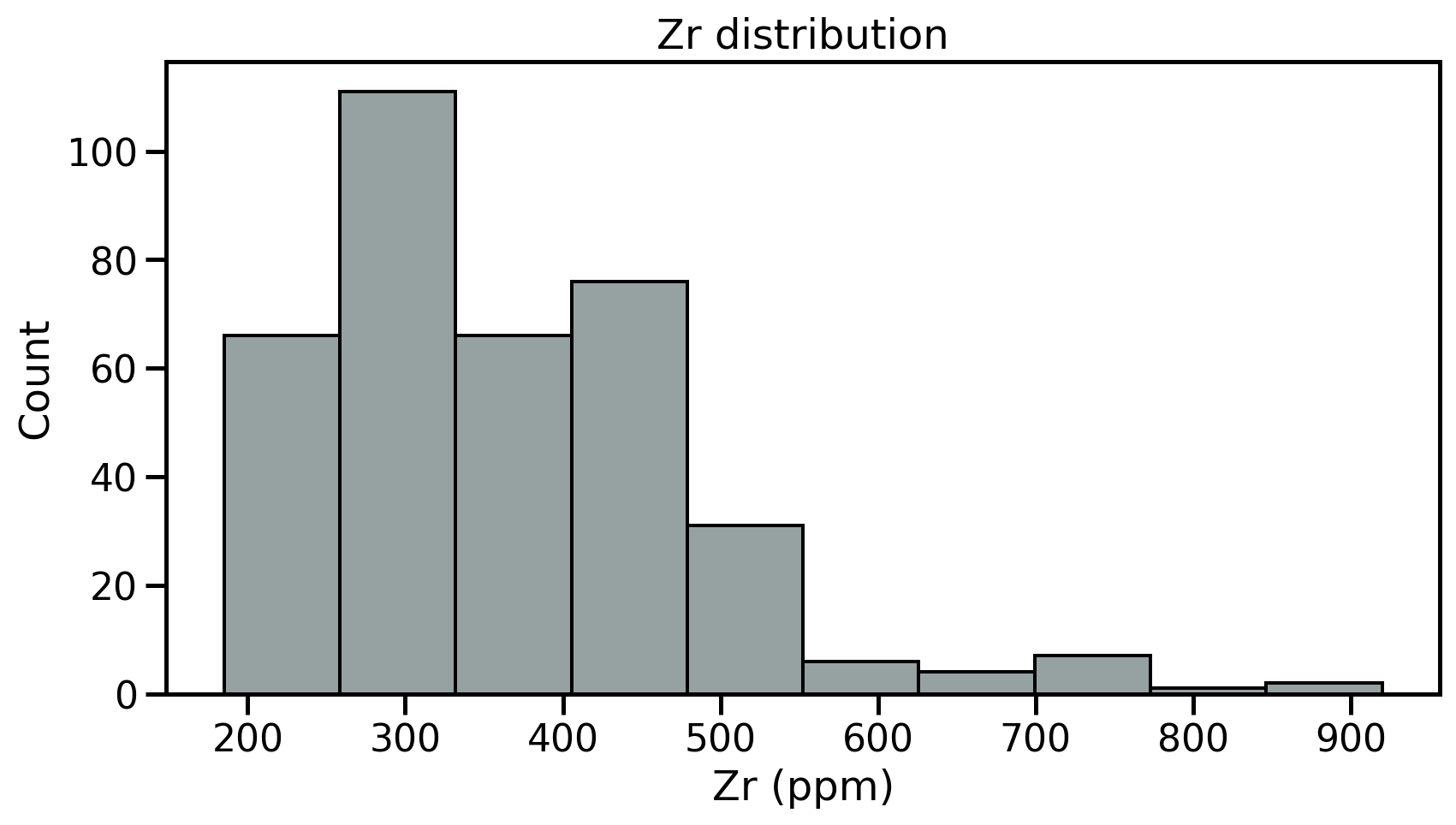
- Fist step: review the samples distribution using histograms
- Divides dataset into equal intervals → bins
- Counts the value in each bin
- Represents the distribution of data
Univariate analyses
- Objective capturing the properties of single variables at the time
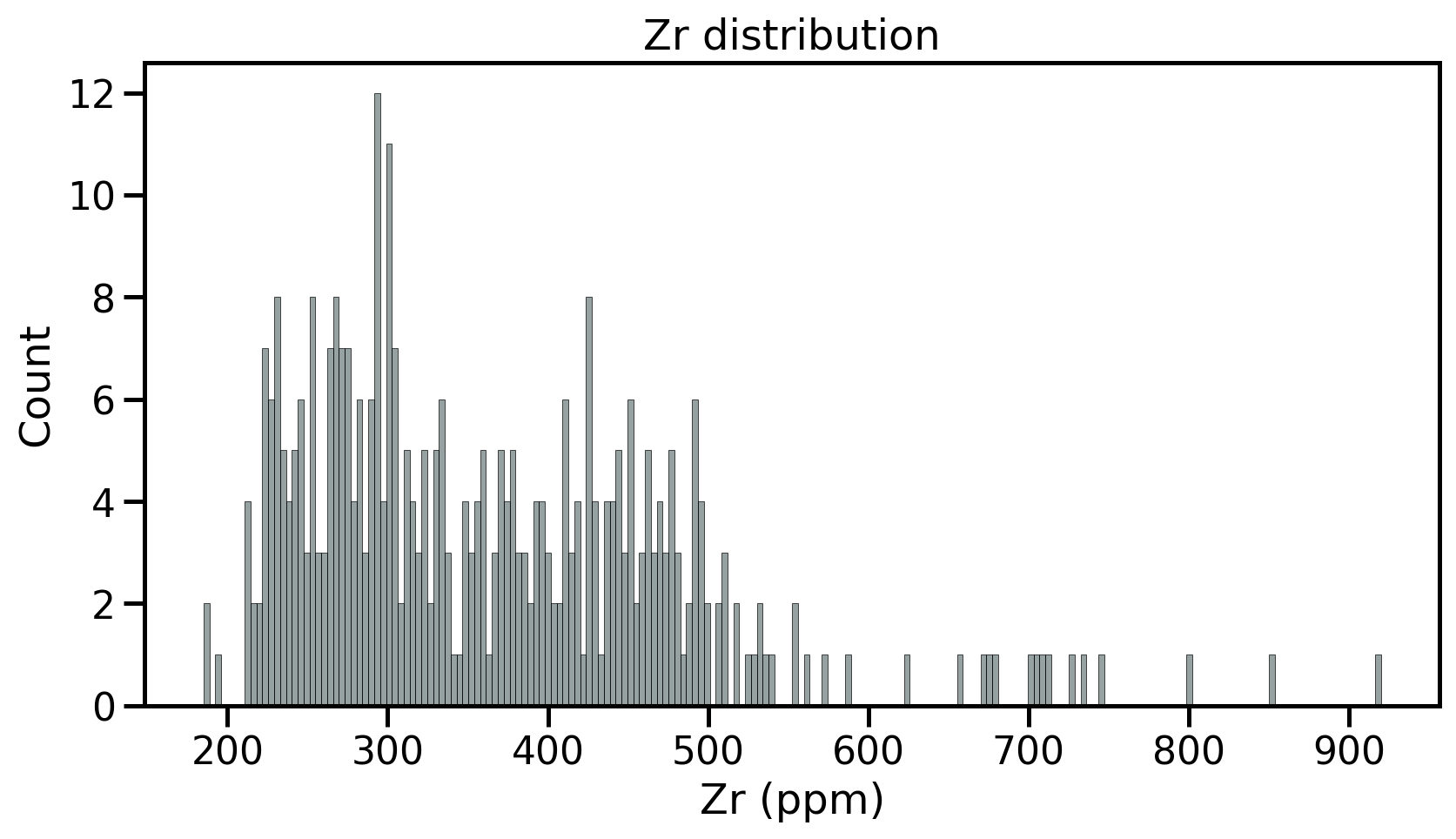
- Fist step: review the samples distribution using histograms
- Divides dataset into equal intervals → bins
- Counts the value in each bin
- Represents the distribution of data
Univariate analyses
- Objective capturing the properties of single variables at the time
- Fist step: review the samples distribution using histograms
- Divides dataset into equal intervals → bins
- Counts the value in each bin
- Represents the distribution of data
Descriptive parameters
Intuitive importance of describing three different characteristics of the distribution:
- The location - or where is the central value(s) of the dataset;
- The dispersion - or how spread out is the distribution of data compared to the central values;
- The skewness - or how symmetrical is the distribution of data compared to the central values;
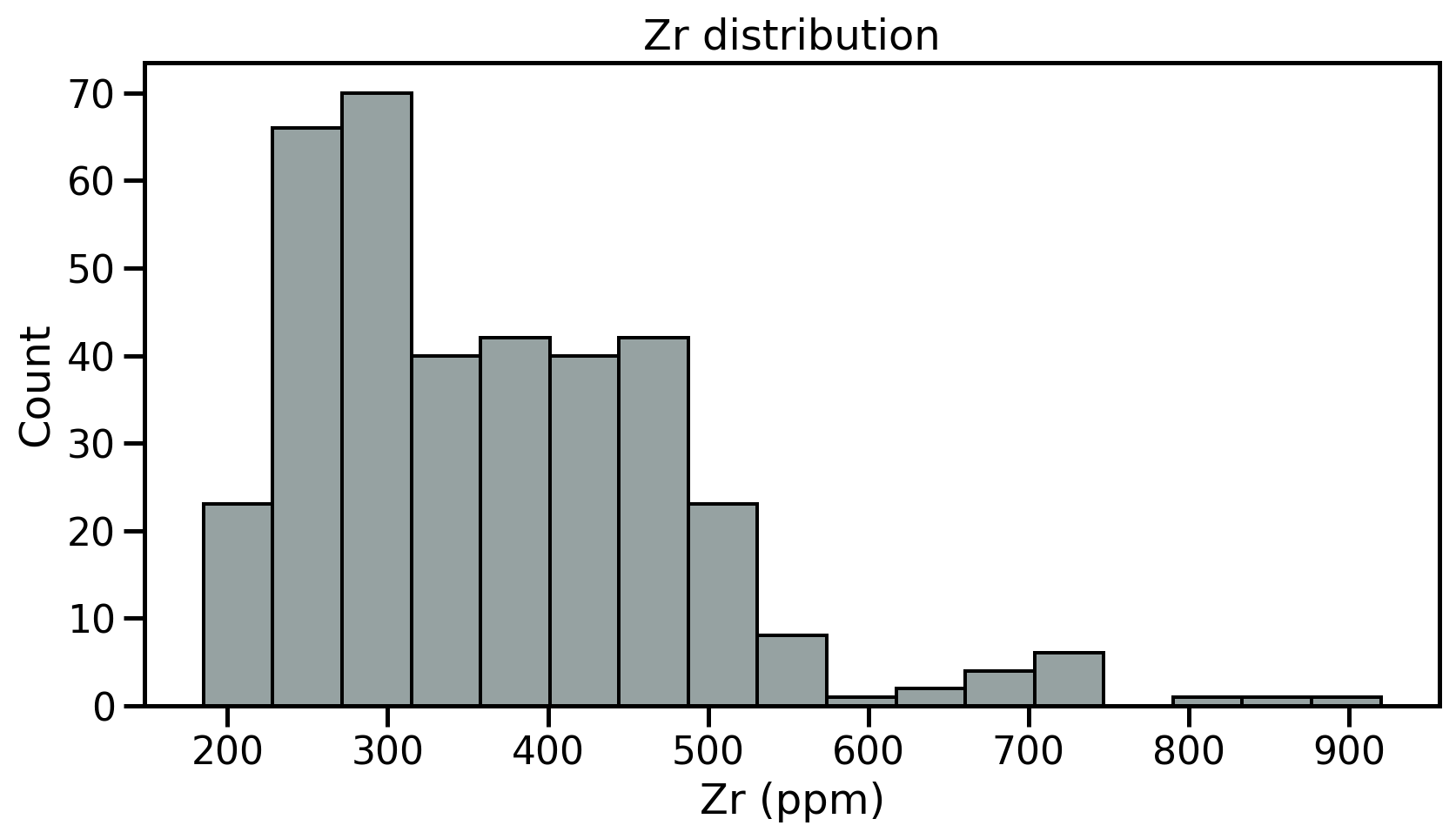
Descriptive parameters
- Ideally → able to constrain full distributions of \(X\)
- theoretical moments
- In practice → only observe a finite sample of \(X\)
- estimators
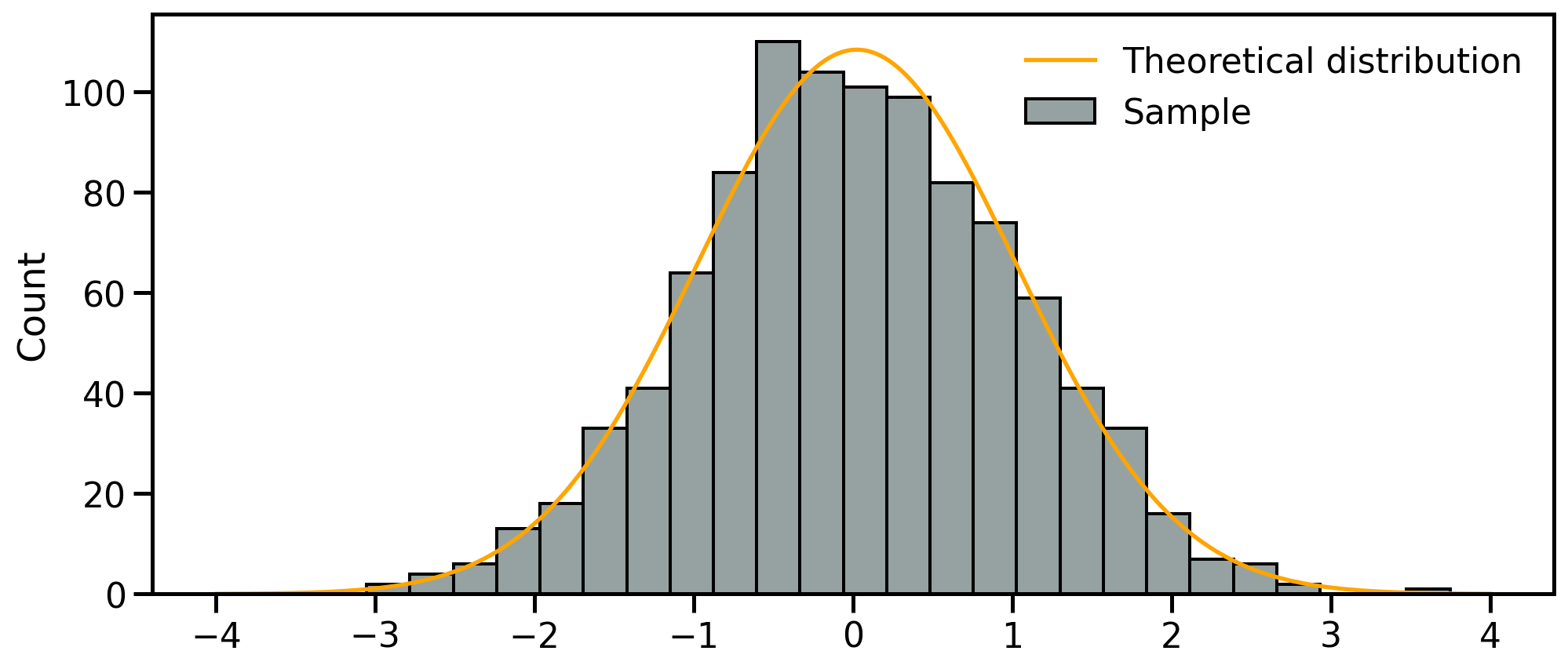
Location I: Mean
The mean is the sum of the values divided by the number of values:
\[ \bar{x} = \frac{1}{n} \sum_{i=1}^{n} x_i \]
- Meaningful to describe symmetrical distributions
- Very sensitive to outliers
df.mean()
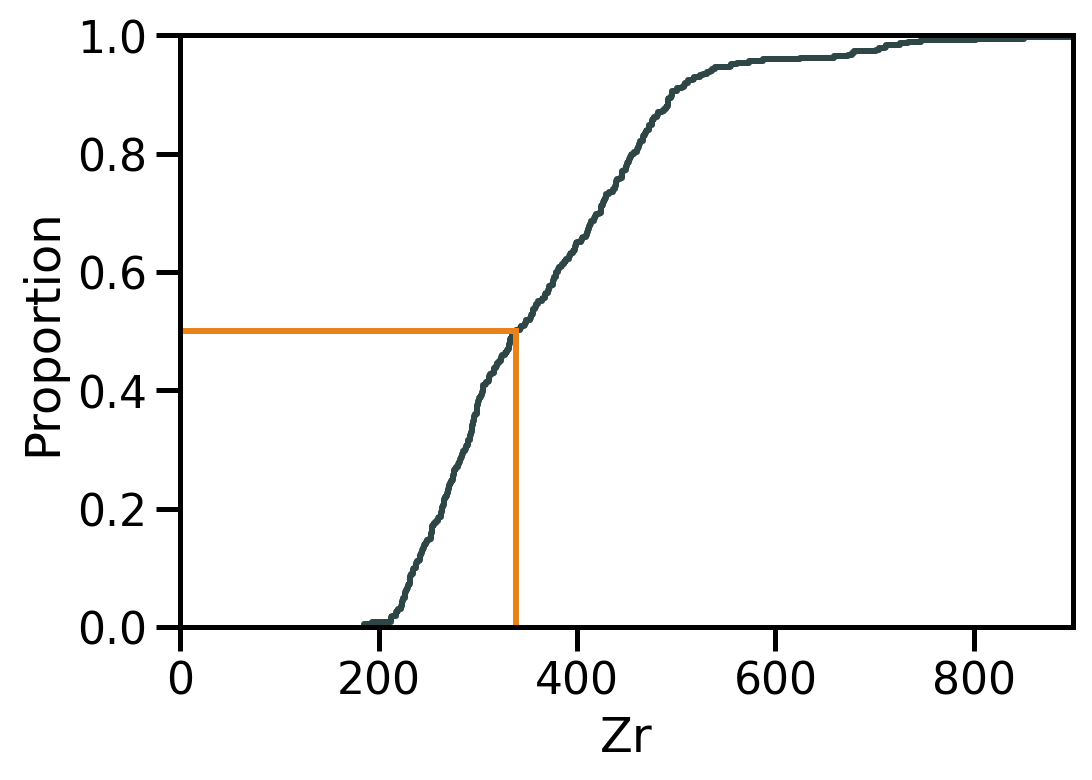
Location II: Median
The median is the value at the exact middle of the dataset.
Location III: Mode
The mode is the value that occur most frequently in a dataset.
- Categorical or discrete numerical data: value(s) that appear most often.
- For continuous data, exact duplicates may be rare
Location: Summary
- Mean: Best for symmetric distributions
- Sensitive to outliers
- Median: Best for skewed or outlier-prone data
- Only based on rank, not magnitude
- Mode: Best for describing the most frequent range in grouped or categorical-like data.
- Otherwise requires binning
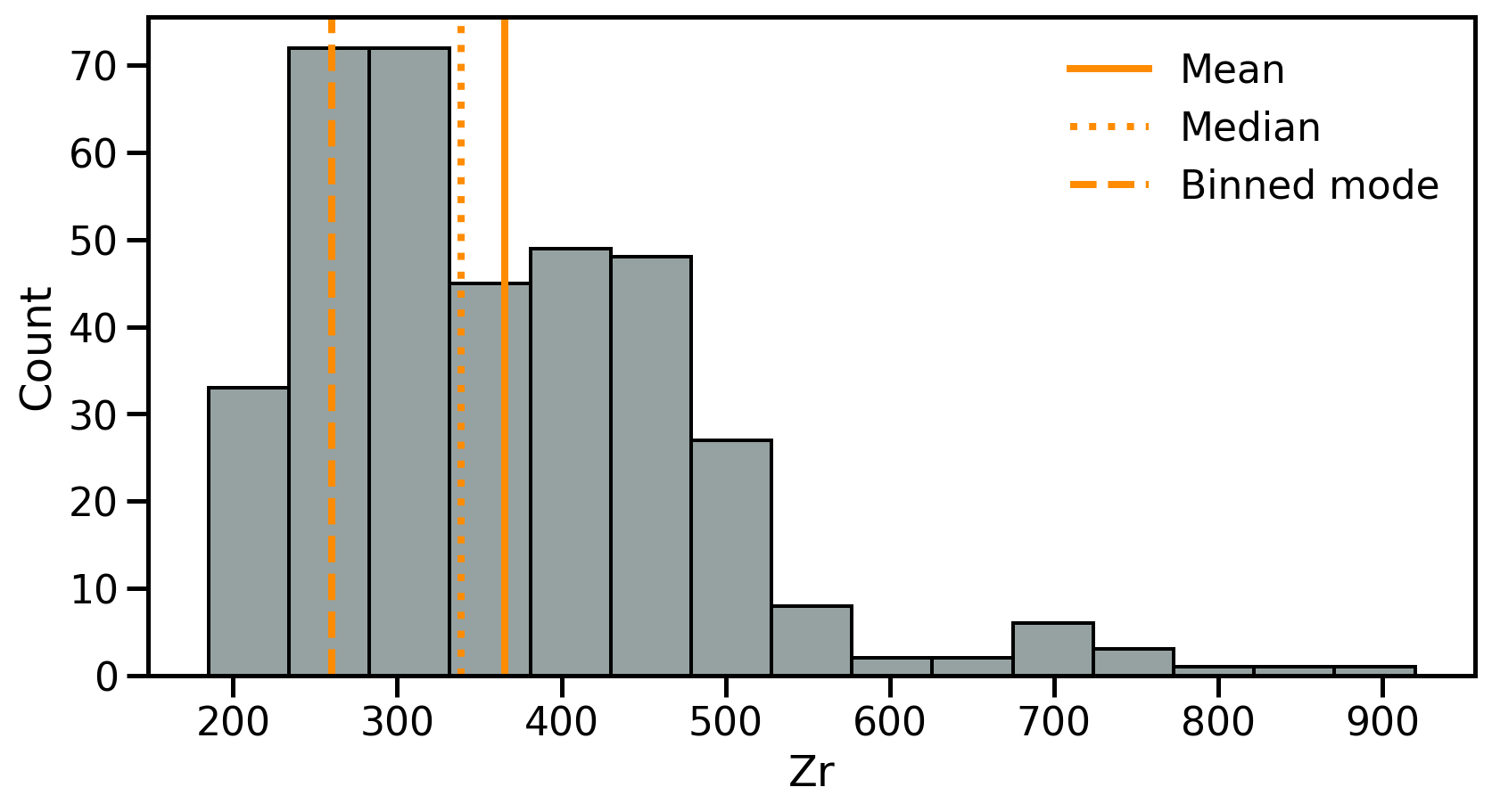
Dispersion 1: Range
The range is the range of value covered in the dataset
- Min →
df.min() - Max →
df.max() - Range:
\[ \text{Range} = \max(x) - \min(x) \]
Don’t use Python reserved keywords as variable names
Names as min, max or range represent native Python variables and cannot be used as variable names!
Dispersion 2: Standard deviation
The sample standard deviation (\(s\)) is the sum of squares differences between data points \(x_i\) and the mean \(\bar{x}\) over \(n\) observations: \[ s = \sqrt{\frac{\sum_{i=1}^{n} (x_i - \bar{x})^2}{n-1} } \]
Dispersion 2: Standard deviation
The theoretical standard deviation (\(\sigma\)) is closely related to the normal distribution as:
- \(1\sigma\) → ~68% of the data
- \(2\sigma\) → ~95% of the data
- \(3\sigma\) → ~99.7% of the data
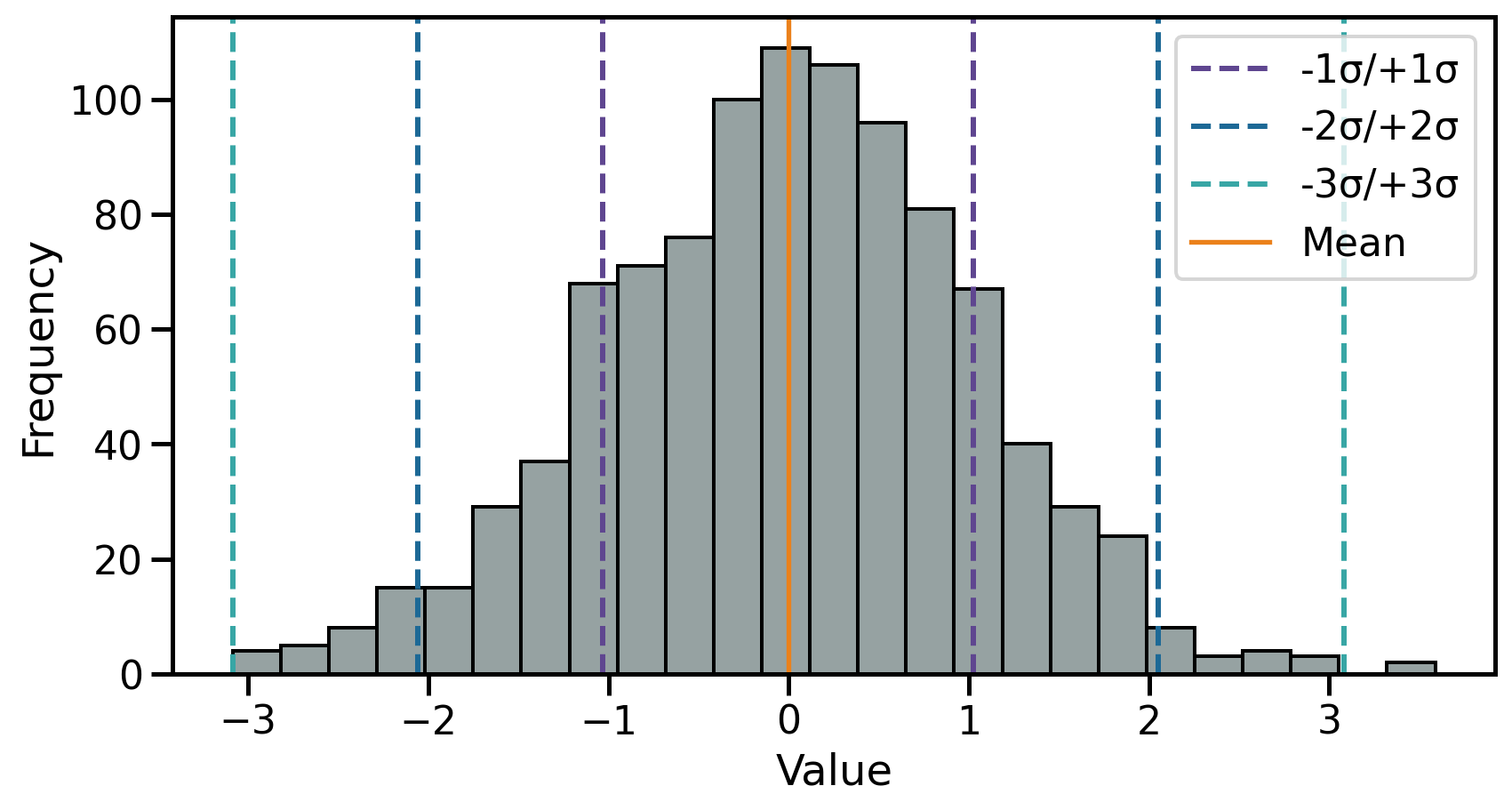
Dispersion 2.1: Variance
The variance (\(s^2\)) is the average of squared deviations from the mean (→ the square of the standard deviation): \[ s^2 = \frac{\sum_{i=1}^{n} (x_i - \bar{x})^2}{n-1} \]
- \(s^2\) is in not the same unit as the data
- Used in modelling (e.g., ANOVA)
df.var()
Rule of thumb
- Comparing data spread → standard deviation
- Statistical modelling → variance
Dispersion 3: Interquartile range
The interquartile range (IQR) indicates how spread out the middle 50% of the data is.
- Difference between the third quartile (Q3) and the first quartile (Q1):
\[ \mathrm{IQR} = Q_3 - Q_1 \qquad(1)\]
- Q1: The value below which 25% of the data fall
- Q3: The value below which 75% of the data fall
Dispersion 3: Interquartile range
| Measure | Number of Divisions | Position(s) in Data / Percentile Equivalent |
|---|---|---|
| Quartiles (Q1, Q2, Q3) | 4 equal parts | Q1 = 25th percentile, Q2 = 50th percentile (median), Q3 = 75th percentile |
| Deciles (D1 … D9) | 10 equal parts | D1 = 10th percentile, D2 = 20th percentile, …, D9 = 90th percentile |
| Percentiles (P1 … P99) | 100 equal parts | P1 = 1st percentile, P2 = 2nd percentile, …, P99 = 99th percentile |
- median =
- 2nd quartile = Q2
- 5th decile = D5
- 50th percentile = P50
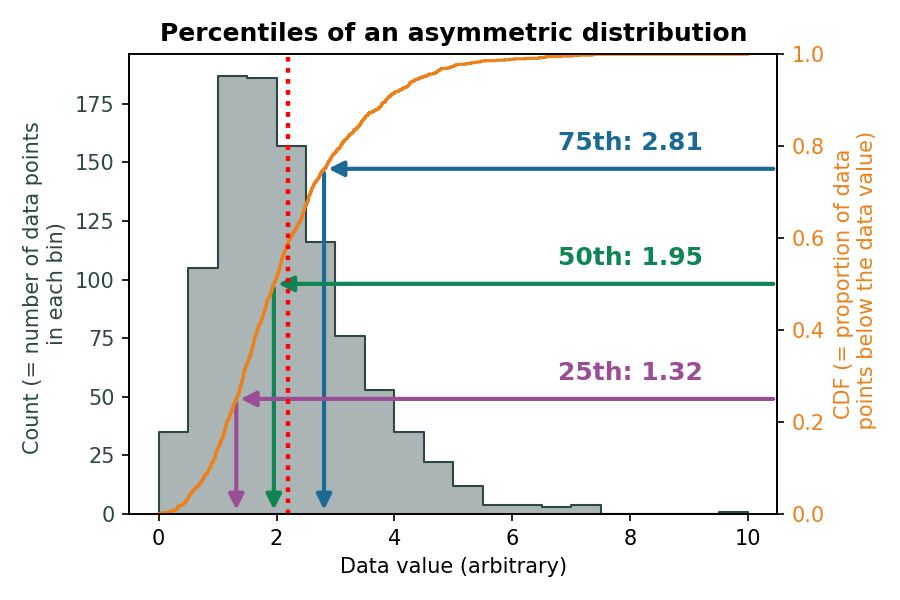
Dispersion 3: Interquartile range
The interquartile range (IQR) indicates how spread out the middle 50% of the data is.
\(Q_1\) and \(Q_3\) for a symmetrical distribution:
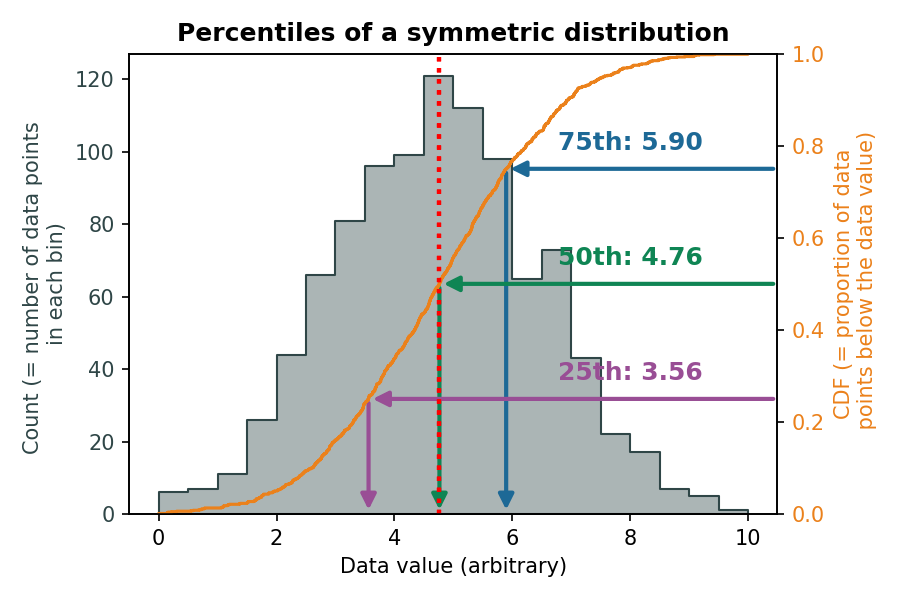
\(Q_1\) and \(Q_3\) for an asymmetrical distribution:
Shape: Skewness
The skewness measures the asymmetry of a distribution and is computed as:
\[ \text{Skewness} = \frac{1}{n} \sum_{i=1}^{n}\left(\frac{x_i-\bar{x}}{s}\right)^{3} \]
In Pandas, the skewness can be computed using .skew()
- 0: Symmetric → e.g., normal distribution
- >0: Righ-skewed → a tail extends to the right
- <0: Left-skewed → a tail extends to the left
Distribution visualisation
Beyond histograms → some plot types report empirical indications of location, dispersion and skewness.
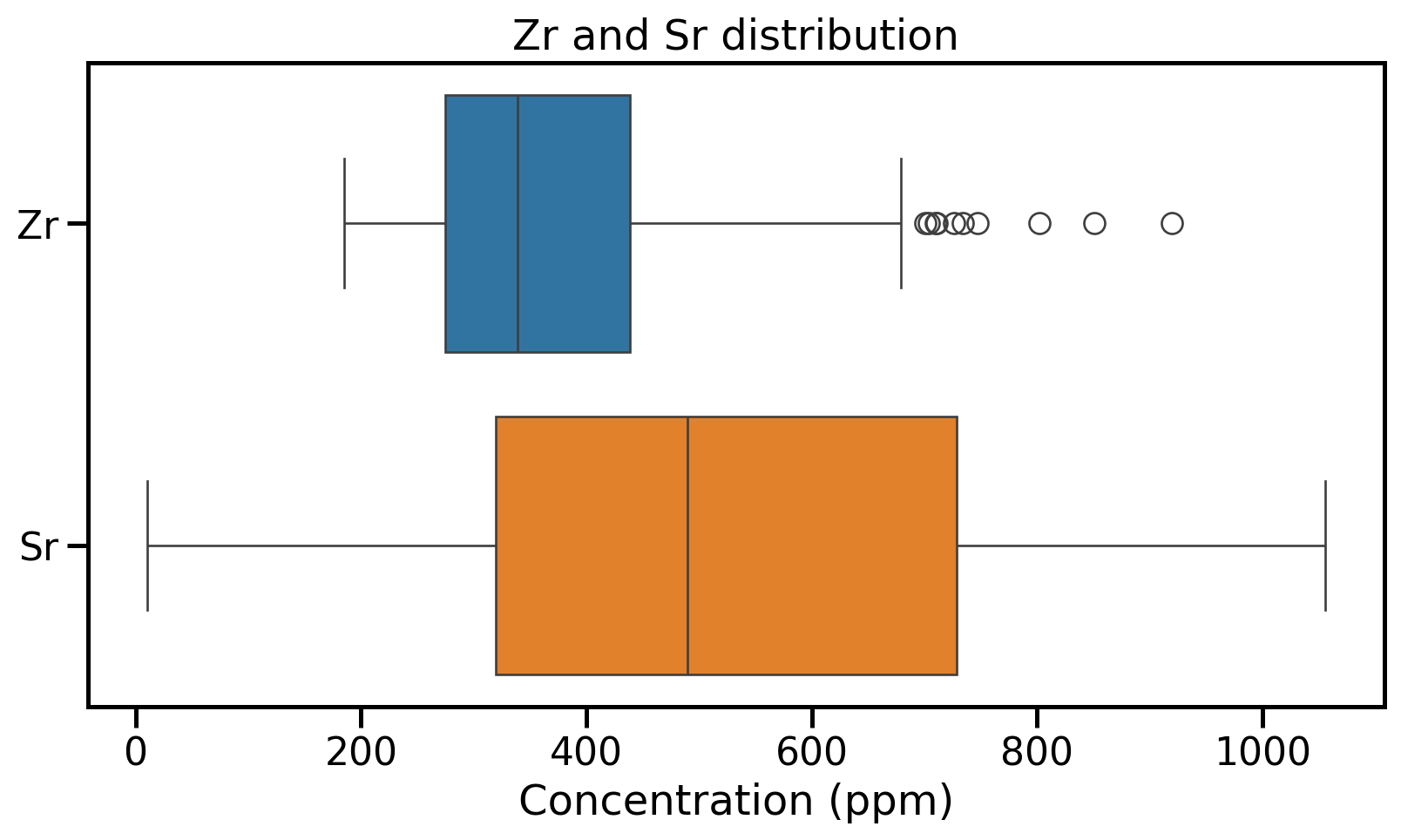
Box and whisker plots
- Box → \(IQR\) + median
- Whiskers 1.5 times the IQR from Q1/Q3
- Outlier → differs from the majority of observations
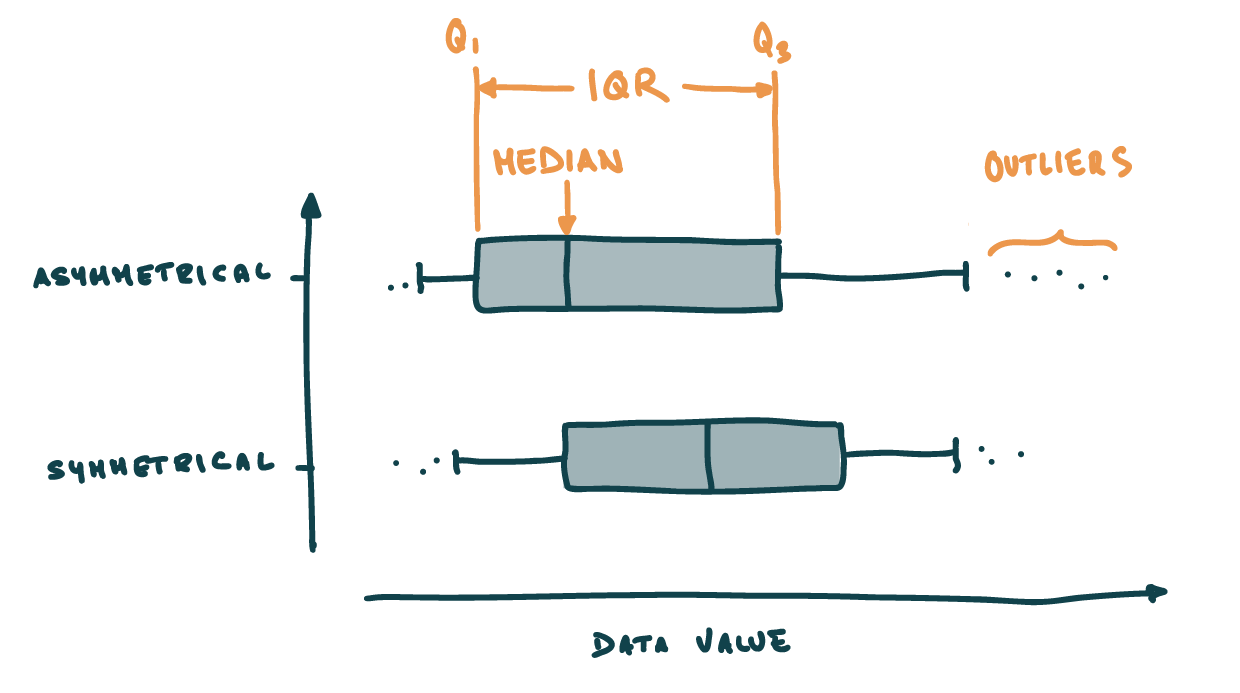
Distribution visualisation: Box plots
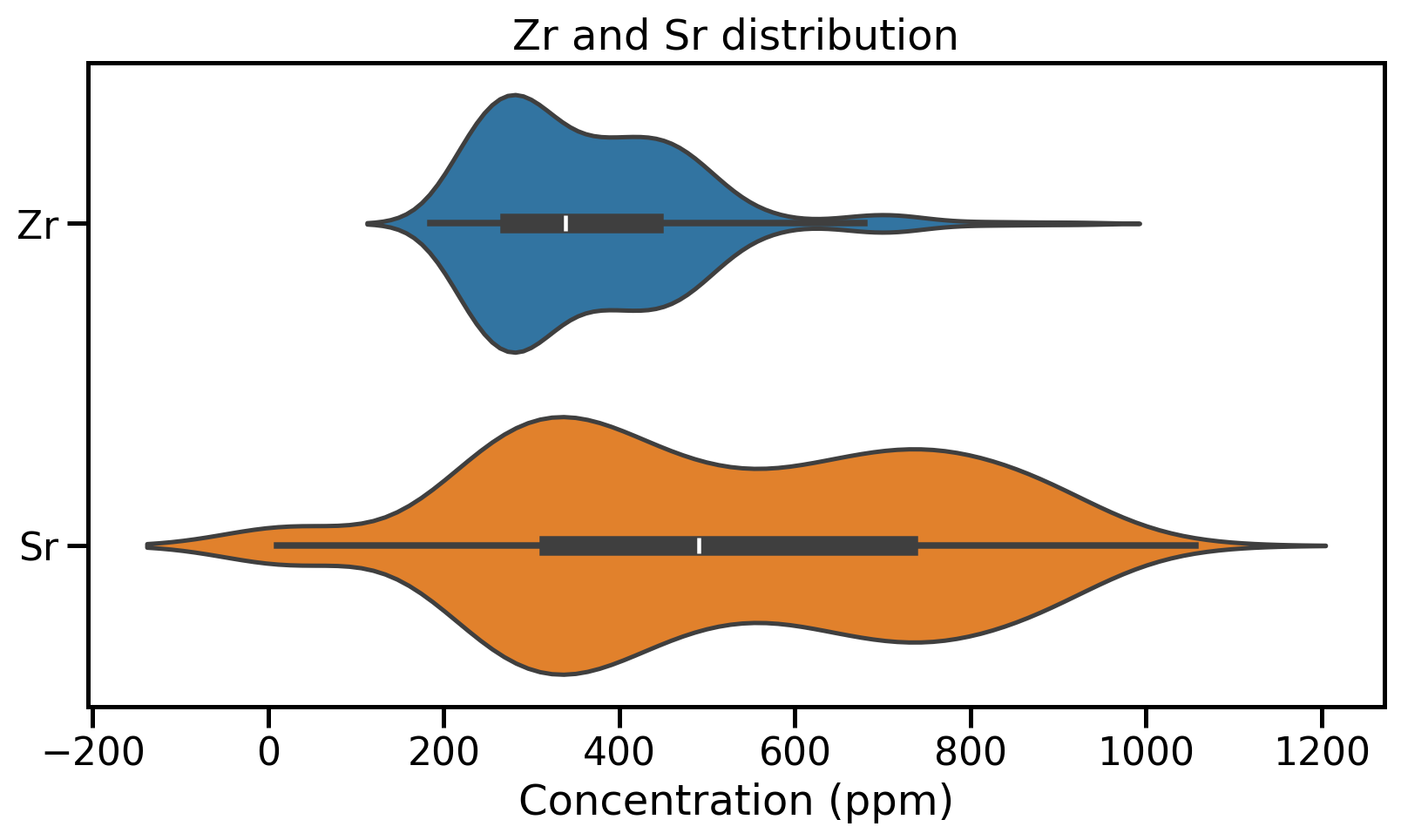
Distribution visualisation: Violin plots
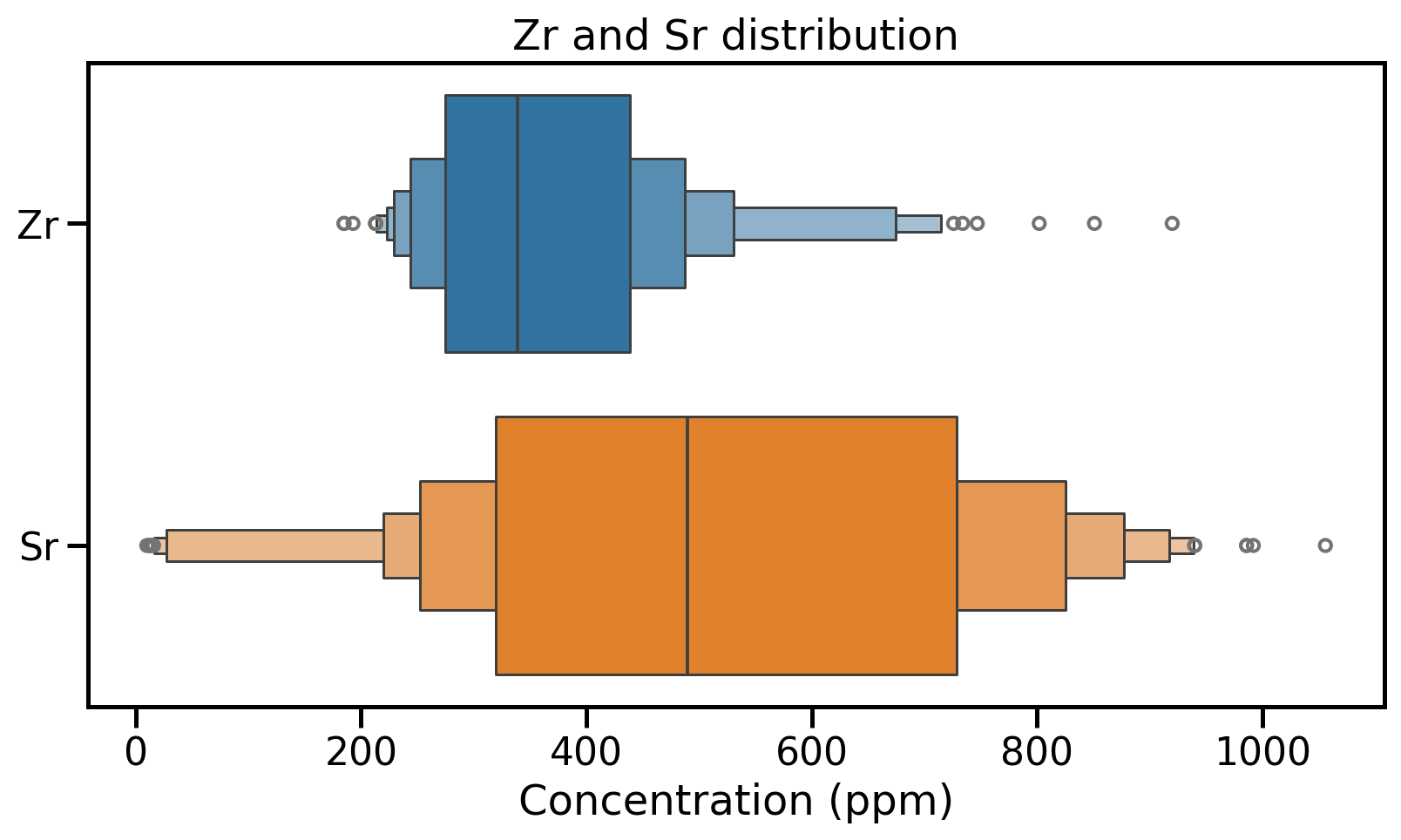
Distribution visualisation: Boxen plots
Your turn!
- Go to https://elste-master.github.io/Data-Science/
- Class 2 > Univariate analyses
- Explore the structure of the glass geochemistry or your dataset
- Get used to the various functions
Bivariate analyses
Bivariate analyses
- Univariate analyses: properties of one variable at the time
- Bivariate analyses: investigate the relationship between two variables
- Visually → pairwise comparison
- Using covariance/correlation → correlation matrix
- First look at linear regression → only visually for now
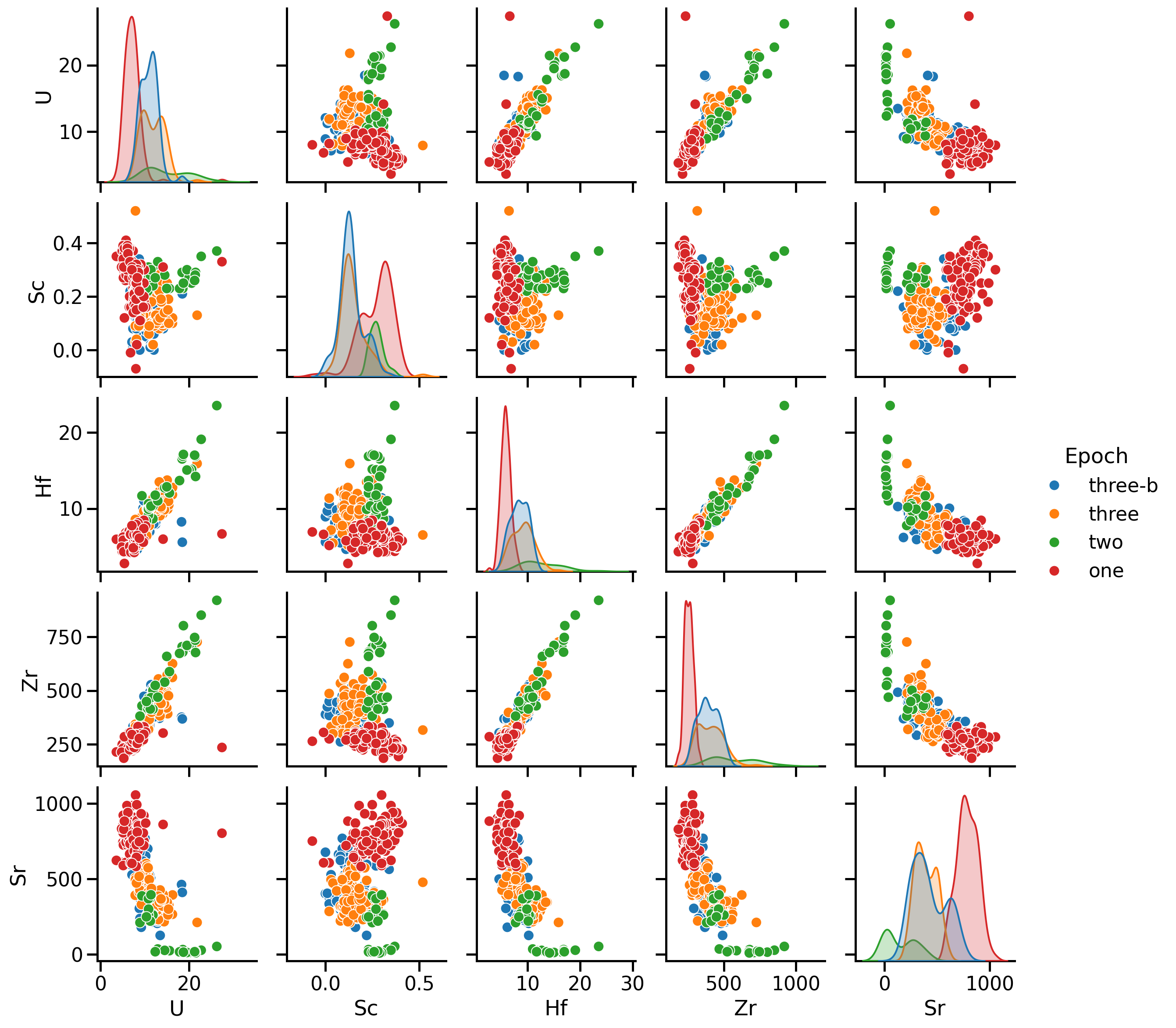
Pairwise comparison
A first visual inspection using sns.pairplot()
- Numerical values
- One categorical value
Covariance
- The covariance captures the direction and magnitude of the linear relationship between the two variables
- The sample covariance is an estimator of the theoretical covariance:
\[ \mathrm{Cov}(X,Y)_{\text{emp}}= \frac{1}{n-1}\sum_{i=1}^n (x_i - \bar{x})(y_i - \bar{y}), \]
- Positive: Increase in \(X\) → increase in \(Y\)
- Negative: Increase in \(X\) → decrease in \(Y\)
Problem: depends on the magnitudes of the variables, so it does not directly reflect the strength of the relationship
Correlation
- The correlation coefficient provides a normalized version of the covariance
- Ranges from \(−1\) to \(1\)
- Indicates both the direction and the strength of the linear relationship
- The sample correlation is computed as:
\[ r(X,Y)_{\text{emp}} = \frac{\mathrm{Cov}(X,Y)_{\text{emp}}}{s_X s_Y} = \frac{\sum_{i=1}^n (x_i - \bar x)(y_i - \bar y)}{\sqrt{\sum_{i=1}^n (x_i - \bar x)^2}\;\sqrt{\sum_{i=1}^n (y_i - \bar y)^2}}, \]
- \(r(X,Y)=-1\)/\(1\) → “perfect” relationship between \(X\) and \(Y\)
- \(r(X,Y)=0\) → independence between \(X\) and \(Y\).
Correlation matrix
- Correlation matrix → square table showing the pairwise correlations between all variables in a dataset
- Suppose we have \(p\) variables \((X_1, X_2, \dots, X_p)\) → the correlation matrix \(\mathbf{C}\) is:
\[ \mathbf{C} = \begin{pmatrix} 1 & r(X_1,X_2) & \dots & r(X_1,X_p) \\ r(X_2,X_1) & 1 & \dots & r(X_2,X_p) \\ \vdots & \vdots & \ddots & \vdots \\ r(X_p,X_1) & r(X_p,X_2) & \dots & 1 \end{pmatrix}. \]
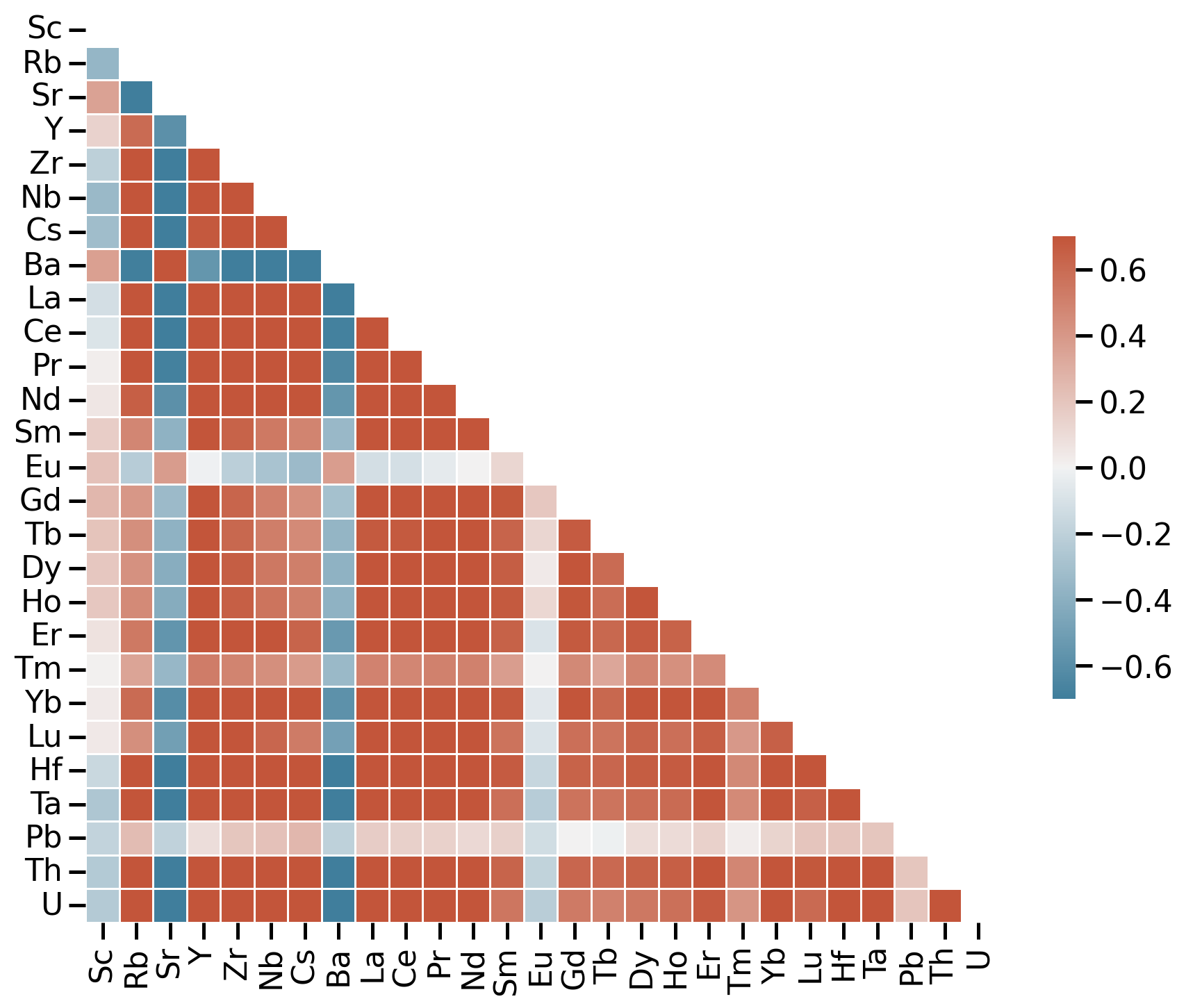
Correlation matrix
Correlation matrix for trace elements
- Use sample correlation coefficient \(r\)
- Red: positive relationship
- Blue: negative relationship
- White: no relationship
Linear regression
- Correlation coefficients → measure of coupling between two continuous variables
- linear regressions attempt modelling this relationship
\[ y = a + bx + \varepsilon \]
where:
- \(y\): dependent (response) variable
- \(x\): independent (predictor) variable
- \(a\): intercept → value of \(y\) when \(x=0\)
- \(b\): slope → how much \(y\) changes by unit of \(x\)
- \(\varepsilon\): error term
→ Objective: Estimate the values of \(a\) and \(b\) that best fit the observed data
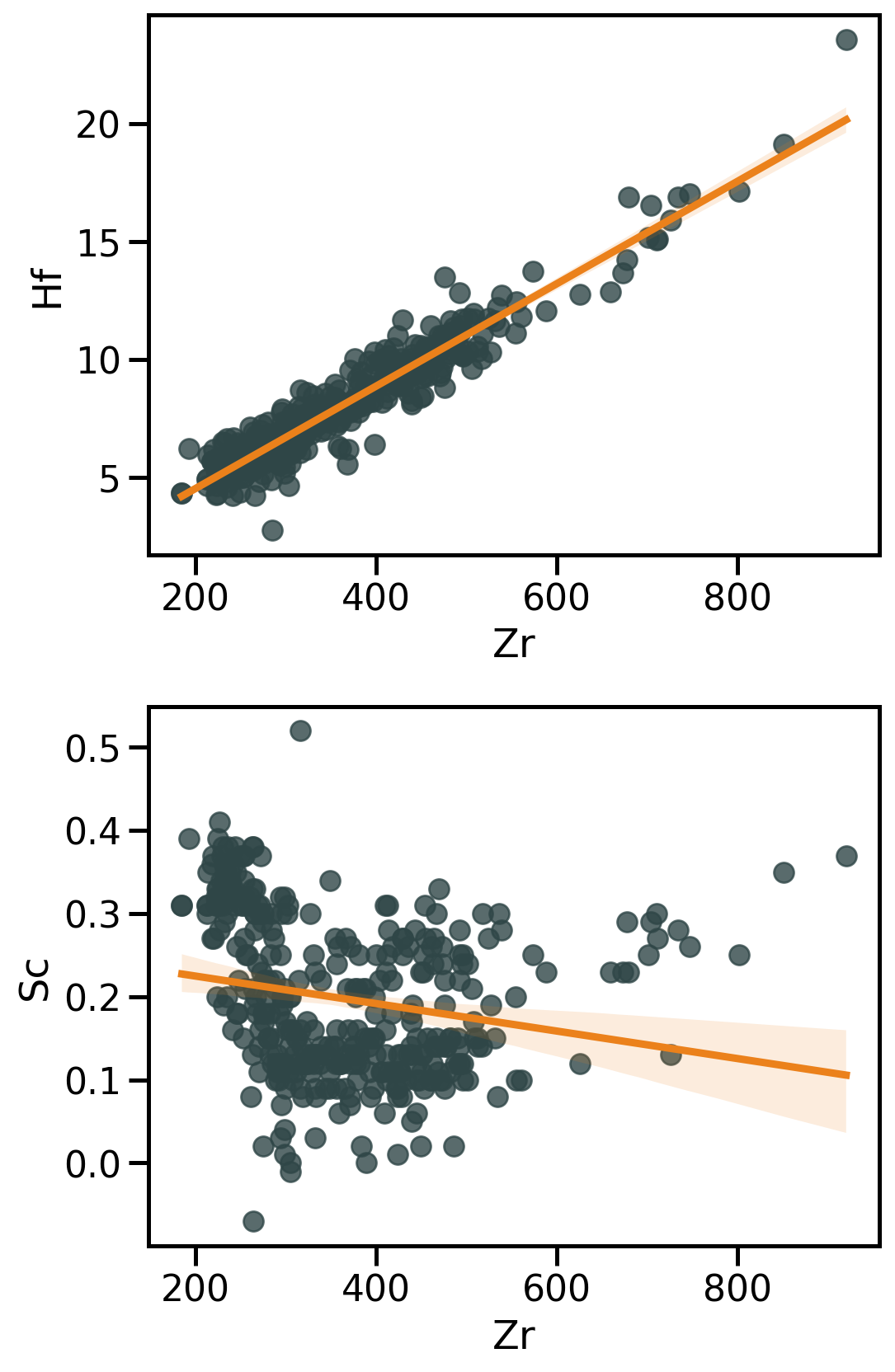
Linear regression with Seaborn
Seaborn has high level functions to visualise regressions → sns.regplot() that produce:
- A scatter plot;
- A linear regression model fit;
- A confidence interval
→ Be critical
→ Further investigations require dedicated stats packages
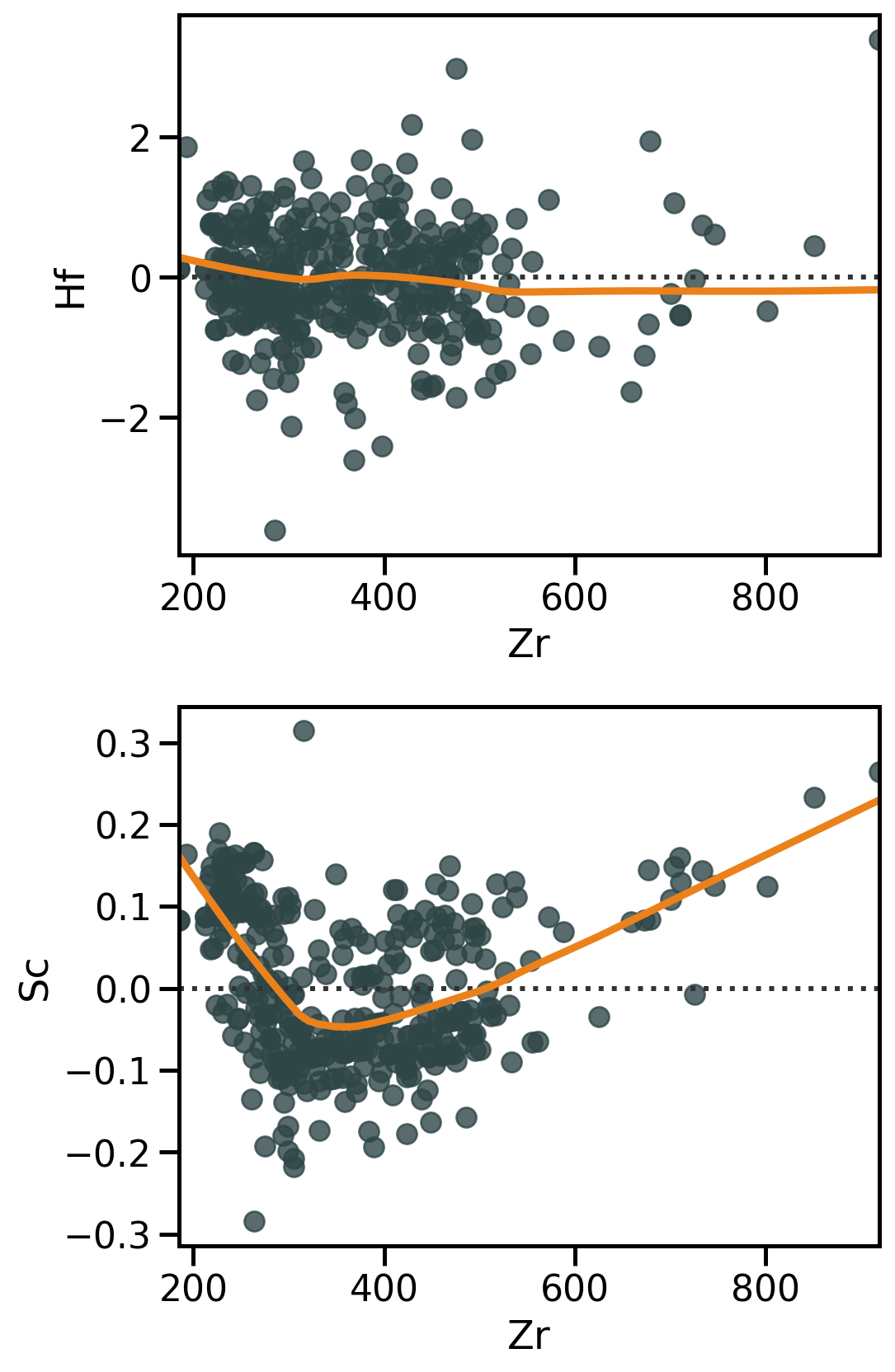
Residual analysis with Seaborn
Linear regression → do residuals show any structure?? → sns.residplot()
Residuals are necessary to:
- Diagnose model fit (→ how much unexplained variation remains);
- Detect patterns that indicate problems (non-linearity, heteroscedasticity, outliers).
Any type of structure in residuals might reveal a violation of linear regression assumptions.
Your turn!
- Go to https://elste-master.github.io/Data-Science/
- Class 2 > Bivariate analyses
- Explore the structure of the glass geochemistry or your dataset
- Get used to the various functions